 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013611035.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 995aa MW: 111067 Da PI: 6.0968 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 187.7 | 1.2e-58 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
l+e ++rwl++ ei++iL n++k+++++e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+++vl+cyYah+e+n++fq
XP_013611035.1 19 LSEaQHRWLRPAEICEILRNYQKFHIASEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSIDVLHCYYAHGEDNENFQ 113
45559****************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rrcyw+Le+el +iv+vhylevk
XP_013611035.1 114 RRCYWILEQELMHIVFVHYLEVK 136
********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 85.888 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.1E-85 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.4E-51 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| SuperFamily | SSF81296 | 3.36E-12 | 402 | 486 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:1.25.40.20 | 1.3E-18 | 583 | 695 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 2.18E-19 | 583 | 694 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 20.208 | 583 | 665 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 2.9E-8 | 584 | 659 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.47E-16 | 589 | 692 | No hit | No description |
| PROSITE profile | PS50088 | 9.458 | 600 | 632 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 1000 | 600 | 629 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 11.22 | 633 | 665 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0016 | 633 | 662 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.32 | 810 | 832 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 9.29E-8 | 811 | 860 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 7.858 | 812 | 840 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0019 | 812 | 831 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0092 | 833 | 855 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.304 | 834 | 858 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 4.4E-4 | 836 | 855 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 995 aa Download sequence Send to blast |
MADRGSFGFA PQLDIQQLLS EAQHRWLRPA EICEILRNYQ KFHIASEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSIDVLH CYYAHGEDNE NFQRRCYWIL 120 EQELMHIVFV HYLEVKGNRI SSSGIKETNS NSLTGSTSVN IDSTANTSST LSPLCEDADS 180 GNRDGWIHGN RVKESDSQRL VGVPALDASF ENPLARYQNL PYNPLLTQTN PSNAGLMSVE 240 GHLRSPLQNQ VNWQIPVQDS LPLQKWPMDS HGTDLALHEN FGTFSSLIGS QNQQQQPIGG 300 SSFQAPFTSV EAAYIPKFGP EDLLYEASAN QTLPLRKSLL KKEDSLKKVD SFSRWVSNEL 360 AEMEDLQMQS SSGGIGWTSV ETAAAASSLS PSLSKDQRFT MIDFWPKWTQ TDTAVEVMVI 420 GTFLLSPEEV TSYSWACMFG EVEVPAEILV DGVLCCHAPP HEVGQVPFYI TCSDRFSCSE 480 VREFDFLPGS ARKLNTADIY GAYTNEASLH VRFENLLARV SSAQEHNVFE DVGEKRRKIS 540 RIMLLKDEKE SFLTSTVEKD LTAVEAKERL VREEFEDKLY LWLIHKVTEE GKGPNILDED 600 GQGVLHLAAA LGYDWAIKPI LAAGVSINFR DANGWSALHW AAFSGREDTV ALLVSLGADS 660 GAVTDPSPEL PLGKTASDLA YGNGHRGISG FLAESSLTSY LEKLTVDGKE DSSTDSSRAK 720 AVQTVAERTA TPMSYGDVPE TLSMKDSLTA VLNATQAADR LHQVFRMQSF QRKQLSEIGD 780 KNEFGLSDEL AVSFAAGKSK KAGGHSSGAA VHAAAVQIQK KYRGWKKRKE FLLIRQRIVK 840 IQAHVRGHQV RKQYRAIIWS VGLLEKIILR WRRKGSGLRG FKRDAVTKAP EPVCAAPAQE 900 DDYDFLKEGR KQTEERLQKA LTRVKSMAQY PEARAQYRRL LTVVEGIREN EASSSSAMNN 960 NNNNTEEAAN YNEEDDLIDI DSLLDDDTFM SLAFE |
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, stems, old leaves, petals, sepals, top of carpels, stigmas, stamen filaments, anthers and siliques, but not in pollen. {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:14581622}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
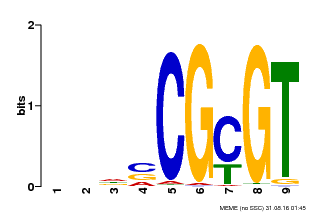
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013611035.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013611035.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A0D3E250 | 0.0 | A0A0D3E250_BRAOL; Uncharacterized protein | ||||
| STRING | Bo9g018010.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4082 | 24 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



