 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013603127.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 292aa MW: 33882.9 Da PI: 5.6429 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 162.1 | 2e-50 | 6 | 140 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.k.kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevl 93
lppGfrFhPtdeelv +yL+++ eg ++el e+i+ +d+yk++Pw+Lp k + +++ ew+fF++rd+ky++g+r nratksgyWkatgkd++++
XP_013603127.1 6 LPPGFRFHPTDEELVGYYLHRRNEGLEIEL-EIIPLMDLYKFDPWELPeKsFLPNRDMEWFFFCHRDRKYQNGSRINRATKSGYWKATGKDRKIV 99
79****************************.99**************95334556888************************************* PP
NAM 94 sk......kgelvglkktLvfykgrapkgektdWvmheyrl 128
++ +++ +g++ktLvfy grap g +t+W mheyrl
XP_013603127.1 100 CQssssssTSSIIGCRKTLVFYLGRAPFGGRTEWAMHEYRL 140
*96666555555***************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.83E-57 | 3 | 164 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 55.75 | 6 | 164 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.5E-27 | 7 | 140 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 292 aa Download sequence Send to blast |
MVRSYLPPGF RFHPTDEELV GYYLHRRNEG LEIELEIIPL MDLYKFDPWE LPEKSFLPNR 60 DMEWFFFCHR DRKYQNGSRI NRATKSGYWK ATGKDRKIVC QSSSSSSTSS IIGCRKTLVF 120 YLGRAPFGGR TEWAMHEYRL FDNDTSQGSL SYKGDFALCR VIKRNEVTLK KCETSSLEVP 180 DEPLTNNVDI PCEARYIGKG SCDASNTLRA SPNFILGSST KGNSQSQTEE DSGFQEFSLP 240 EIEYPPEFFA DLNFDWGMEN PFPFVQYQEA HVNNEVINYD VLRNHHCYHI FT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 7e-49 | 6 | 170 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 7e-49 | 6 | 170 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 7e-49 | 6 | 170 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 7e-49 | 6 | 170 | 17 | 171 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 7e-49 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swm_B | 7e-49 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swm_C | 7e-49 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swm_D | 7e-49 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swp_A | 7e-49 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swp_B | 7e-49 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swp_C | 7e-49 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 3swp_D | 7e-49 | 6 | 170 | 20 | 174 | NAC domain-containing protein 19 |
| 4dul_A | 7e-49 | 6 | 170 | 17 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 7e-49 | 6 | 170 | 17 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, rosettes leaves, cauline leaves and stems. {ECO:0000269|PubMed:24285786}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in the positive regulation of abscisic acid (ABA) responsive genes. Acts as a positive factor of ABA-mediated responses. Involved in the transcriptional activation of ABA-inducible genes in response to dehydration and osmotic stresses. Plays a positive role in both stomatal closure and water loss under dehydration stress conditions. Acts synergistically with ABF2 to activate the dehydration stress-response factor RD29A transcription. Binds to the consensus core cis-acting elements 5'-CGTA-3' and 5'-CACG-3' at the RD29A promoter (PubMed:24285786). Involved in hypocotyl graft union formation. Required for the auxin-mediated promotion of vascular tissue proliferation during hypocotyl graft attachment (PubMed:25182467). {ECO:0000269|PubMed:24285786, ECO:0000269|PubMed:25182467}. | |||||
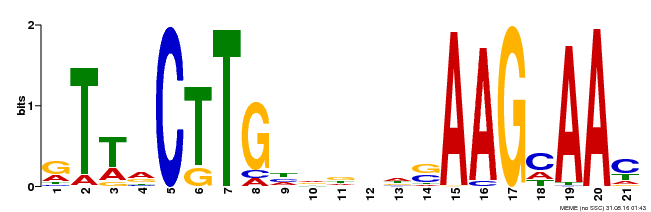
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00175 | DAP | Transfer from AT1G32510 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013603127.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by abscisic acid (ABA), dehydration and osmotic stress. {ECO:0000269|PubMed:24285786}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP002684 | 1e-105 | CP002684.1 Arabidopsis thaliana chromosome 1 sequence. | |||
| GenBank | F5D14 | 1e-105 | AC007767.3 Sequence of BAC F5D14 from Arabidopsis thaliana chromosome 1, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013603127.1 | 0.0 | PREDICTED: NAC domain-containing protein 86 isoform X2 | ||||
| Swissprot | Q9LS24 | 3e-85 | NAC96_ARATH; NAC domain-containing protein 96 | ||||
| TrEMBL | A0A0D3DKT5 | 0.0 | A0A0D3DKT5_BRAOL; Uncharacterized protein | ||||
| STRING | Bo8g028730.1 | 0.0 | (Brassica oleracea) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G32510.1 | 1e-154 | NAC domain containing protein 11 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



