 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_013591899.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 817aa MW: 94349.2 Da PI: 7.6889 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 89.5 | 4.6e-28 | 62 | 150 | 2 | 91 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
fY++YAk++GF++++++s++sk+++ +++++f Cs++g++ e+++ +++ r++++++t+Cka+++vk+++dg+w ++++ +eHnHel p
XP_013591899.1 62 FYQDYAKSMGFTTSIKNSRRSKKTKDFIDAKFACSRYGSTPESESG-SSSGRRSSVKKTDCKASMHVKRKSDGRWIIHEFIKEHNHELLP 150
****************************************999998.888889999*******************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.1E-25 | 62 | 150 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 2.0E-25 | 271 | 364 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 8.716 | 551 | 587 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 7.2E-6 | 553 | 586 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 2.1E-5 | 562 | 589 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 817 aa Download sequence Send to blast |
MDLQESNNNQ MSDANMVESH NNNEDVGGVI VDDLRTGGHL GALDLEPRDG IAFDTHEAAY 60 TFYQDYAKSM GFTTSIKNSR RSKKTKDFID AKFACSRYGS TPESESGSSS GRRSSVKKTD 120 CKASMHVKRK SDGRWIIHEF IKEHNHELLP ALAYHFRIQR NIKLAEKNNI DILHAVSERT 180 RKMYVEMSRQ SGGYKNNKGL LLLQTSEVDK GRRCLALEEG ESQLLLEYFK RIKKENPRFF 240 YAIDLNEEQR LRNLFWADAK SRDDYVSFND VVSFDTTYVK FNDKLPLALF IGVNHHSQPM 300 LLGCGLVADE STETFAWLIK TWLRAMGARA PKVILTDQDK LLMAAVSELL PNTRHCFALW 360 HVLEKLPEYF SHVMKRHENF LRKFNKCIFR SWTSDQFDMR WWKMVSQFGL ENDEWLLWLH 420 EHRQKWVPTF MSEVFLAGMS TSQRAESVNS FFDKYVHKKI TLKEFLRLYG VMLQNRYEEE 480 SVADFDTCHK QPALKSPSPW EKQMAATYTH TIFKKFQVEV LGVVACHPRK EKEDENMVTF 540 RVQDCEKDDE FLVTWSKTQS ELCCFCYMFE YKGFLCRHAL MILQMCGFAS VPPRYILKRW 600 TKDAKSGGAD QIQTRVQRYN DLCCRAIELS EEGCVSEENY NIVLRTLVET LKNCVDLNHA 660 RNNFAESNSQ LNNGAHEEEN HVLAAVKATK KKTVVRKRKG QPEASQMLES QPSLQPMENI 720 SSEGMSMSGY YGPQQNVQGL LNLMEPPHEG YYVNQRTIQG LGQLNSIAPV QDSFFTNQQA 780 MPGLGQMDFR PPPNFAYNLQ DEHLRSAQLP GSSSRQL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
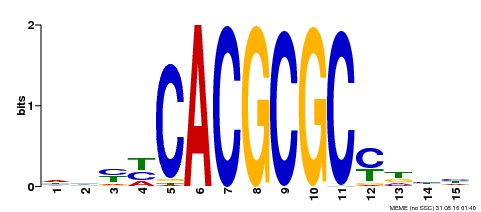
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_013591899.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY078941 | 0.0 | AY078941.1 Arabidopsis thaliana AT4g15090/dl3590w mRNA, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013591899.1 | 0.0 | PREDICTED: protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Refseq | XP_013702274.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1-like | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A0D3CSJ4 | 0.0 | A0A0D3CSJ4_BRAOL; Uncharacterized protein | ||||
| TrEMBL | A0A3P6G8H5 | 0.0 | A0A3P6G8H5_BRAOL; Uncharacterized protein | ||||
| STRING | Bo6g054750.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||



