 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_011091721.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Pedaliaceae; Sesamum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 828aa MW: 90414.9 Da PI: 6.3006 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.1 | 4.6e-21 | 131 | 186 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
XP_011091721.1 131 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 186
688999***********************************************999 PP
| |||||||
| 2 | START | 205.9 | 1.6e-64 | 337 | 562 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetl 82
ela++a++elvk+a+ +ep+W +s e++n +e+l++f++ + + +ea+r++g+v+ ++ lve+l+d++ +W e+++ + +t+
XP_011091721.1 337 ELALAAMDELVKMAQTDEPLWLRSLeggrEILNHEEYLRTFTPCIGmkpngFVTEASRETGMVIINSLALVETLMDSN-KWAEMFPciiaRTSTT 430
5899**************************************9988999*9***************************.**************** PP
EEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEECTCEEEEEEEE- CS
START 83 evissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepksnghskvtwveh 169
+vissg galqlm+a+lq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++ + + +++lpSg+++++++ng+skvtwveh
XP_011091721.1 431 DVISSGmggtrnGALQLMNAALQVLSPLVPvREVNFLRFCKQHAEGVWAVVDVSIDTIRETSGgPTTFPNCRRLPSGCVVQDMPNGYSKVTWVEH 525
********************************************************9999998888999************************** PP
EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 170 vdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
v+++++++h+l+r+l++ g+ +ga++wvatlqrqce+
XP_011091721.1 526 VEYDESVVHQLYRPLISAGMGFGAQRWVATLQRQCEC 562
***********************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.24E-20 | 114 | 188 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-21 | 118 | 188 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.216 | 128 | 188 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.9E-18 | 129 | 192 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.34E-18 | 130 | 188 | No hit | No description |
| Pfam | PF00046 | 1.4E-18 | 131 | 186 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 163 | 186 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.74 | 328 | 565 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 6.87E-32 | 330 | 562 | No hit | No description |
| CDD | cd08875 | 2.34E-124 | 332 | 561 | No hit | No description |
| Pfam | PF01852 | 3.8E-56 | 337 | 562 | IPR002913 | START domain |
| SMART | SM00234 | 7.2E-51 | 337 | 562 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.47E-24 | 589 | 821 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 828 aa Download sequence Send to blast |
MNFGDFLDNN SCGGGGARIV SDIPYSNSNS NNNNAITSDI NSMPTGAIAH PRLVSHSLTT 60 KPMFNSPGLS LALQTSMEAQ GDMARMAENY ELSNVGGRRS RDEEHESRSG SDNMDGASGD 120 DQDAADKPPR KKRYHRHTPQ QIQELEALFK ECPHPDEKQR LELSKRLCLE TRQVKFWFQN 180 RRTQMKTQLE RHENSILRQE NDKLRAENLS IREAMRNPIC TNCGGPAIIG EISLEEQHLR 240 IENARLKDEL DRVCALAGKF LGRPISSLAA PMPNSSLELG VGSNGFGGLN TIPSTMPLVV 300 PSDFGMGISS PLPMVTPKAT MNISPIERSL ERSMYLELAL AAMDELVKMA QTDEPLWLRS 360 LEGGREILNH EEYLRTFTPC IGMKPNGFVT EASRETGMVI INSLALVETL MDSNKWAEMF 420 PCIIARTSTT DVISSGMGGT RNGALQLMNA ALQVLSPLVP VREVNFLRFC KQHAEGVWAV 480 VDVSIDTIRE TSGGPTTFPN CRRLPSGCVV QDMPNGYSKV TWVEHVEYDE SVVHQLYRPL 540 ISAGMGFGAQ RWVATLQRQC ECLAILMSST VPVREHTAIT GGGRRSMLKL AQRMTNNFCA 600 GVCASSVHKW NKLRTENVDD DVQVMTRKSV DDPGEPPGIV LSAATSVWLP VSPQRLFDFL 660 RDERLRSEWD ILSNGGPMQE MAHIAKGQDH GNCVSLLRAS AMNSSQSSML ILQETCIDAA 720 GSLVVYAPVD IPAMHVVMNG GDSAYVALLP SGFSIVPDGP GSRGPPSNGP TSNGGATHRV 780 SGSLLTVAFQ ILVNSLPTAK LTVESVETVN NLISCTVQKI KAALHCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
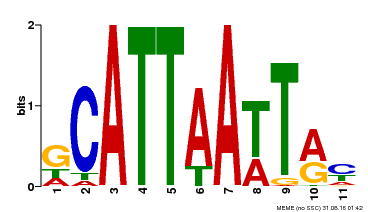
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_011091721.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011091721.2 | 0.0 | LOW QUALITY PROTEIN: homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A2G9HWV6 | 0.0 | A0A2G9HWV6_9LAMI; Transcription factor DLX | ||||
| STRING | VIT_15s0048g02000.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1626 | 24 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



