 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010918971.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 272aa MW: 29795.2 Da PI: 5.419 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 63 | 6.5e-20 | 121 | 172 | 2 | 56 |
AP2 2 gykGVrwdkkrgrWvAeIrd.psengkrkrfslgkfgtaeeAakaaiaarkklege 56
+++GVr+++ +g+++AeIrd + k +r++lg+f taeeAa a+++a+++++g+
XP_010918971.1 121 HFRGVRRRP-WGKFAAEIRDtSR---KGARVWLGTFNTAEEAALAYDEAALRMRGP 172
79*******.**********331...34**************************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.16E-29 | 120 | 179 | No hit | No description |
| Pfam | PF00847 | 1.6E-15 | 121 | 171 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 5.23E-23 | 121 | 180 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 8.2E-32 | 121 | 180 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 23.209 | 121 | 179 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.8E-37 | 121 | 185 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.0E-10 | 122 | 133 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.0E-10 | 145 | 161 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 272 aa Download sequence Send to blast |
MASELDRSET NDKLLQSVWA SFITGGAPSS CSSDESHKKH AASQEWNEMP HLEGGEENLA 60 VLQRLPSLGR WISMGAESWD ALLSETIASN SCETSPLYSG YDACSLSKPK TLAAEKVAPK 120 HFRGVRRRPW GKFAAEIRDT SRKGARVWLG TFNTAEEAAL AYDEAALRMR GPQAHLNFPM 180 EKVTKASEGA HLEQALAGTT SGTACPVDGC IKTNHNPRKR AAREWDTYDE VPAEPPAWKR 240 NSSIEGMGSN QADIVELQDL GIDYLENLLS SF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 1e-27 | 120 | 181 | 4 | 65 | ATERF1 |
| 3gcc_A | 1e-27 | 120 | 181 | 4 | 65 | ATERF1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00441 | DAP | Transfer from AT4G18450 | Download |
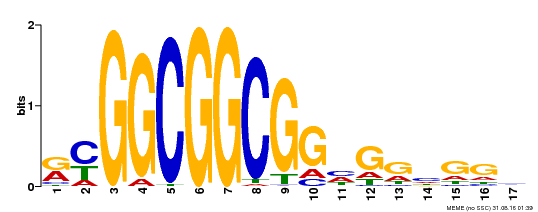
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010918971.1 | 0.0 | ethylene-responsive transcription factor 1B | ||||
| TrEMBL | A0A2H3ZCS4 | 1e-155 | A0A2H3ZCS4_PHODC; ethylene-responsive transcription factor 1B-like | ||||
| STRING | XP_008798293.1 | 1e-145 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP11616 | 27 | 36 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18450.1 | 3e-28 | ERF family protein | ||||



