 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010916598.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 451aa MW: 50703.2 Da PI: 5.239 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 116.6 | 1.5e-36 | 16 | 106 | 2 | 101 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkellek 96
Fl+k+ye+++d+++++++sws +++sf+v+++ ef++++LpkyFkh+nf+SFvRQLn+YgF+k++ ++ weF++++F +g+++ll++
XP_010916598.1 16 FLTKTYEMVDDPSTNSIVSWSPSNSSFIVWNPPEFSRDLLPKYFKHNNFSSFVRQLNTYGFRKIDPDQ---------WEFANEEFIRGQRHLLKN 101
9********************999*****************************************999.........****************** PP
XXXXX CS
HSF_DNA-bind 97 ikrkk 101
i+r+k
XP_010916598.1 102 IHRRK 106
***98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 3.3E-40 | 8 | 100 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SuperFamily | SSF46785 | 1.77E-36 | 11 | 105 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 2.4E-65 | 12 | 105 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 3.5E-32 | 16 | 105 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 6.0E-21 | 16 | 39 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 6.0E-21 | 54 | 66 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 55 | 79 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 6.0E-21 | 67 | 79 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 451 aa Download sequence Send to blast |
MEGSQGGSAS NSPPPFLTKT YEMVDDPSTN SIVSWSPSNS SFIVWNPPEF SRDLLPKYFK 60 HNNFSSFVRQ LNTYGFRKID PDQWEFANEE FIRGQRHLLK NIHRRKPIHS HSLHHQGNSG 120 PLAESERREL EDEIERLKCE KGLLLSELQK HAQQHHGMEQ QMQSLEERLH AMENRQRNMM 180 AFLTQVMQKP GFLSNCVQQS EFHTKKRRLP KANYFCEDSN MEENRIISFQ GMSSEKSDIA 240 SMHALDIEPF EKMESSLNSL ENFFRGVSQA SGEDMYYDSA LPSLPSGVVI MEMNASSGDT 300 DVNLQSPSPK LHPSSPCLGD IHSSPELAES TSYAESPALP ITEIHADIRS KASEIDVNSA 360 PAAPECDSSR DGAAGTAACT MAAGVNDVFW EQFLTETPGS SDTQEVQSER RDANDTRSDG 420 KMGERGNIWW SRKNVEHLAE KMGHLTSAEK T |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 4e-28 | 6 | 105 | 19 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 4e-28 | 6 | 105 | 19 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 4e-28 | 6 | 105 | 19 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA of heat shock promoter elements (HSE). {ECO:0000250}. | |||||
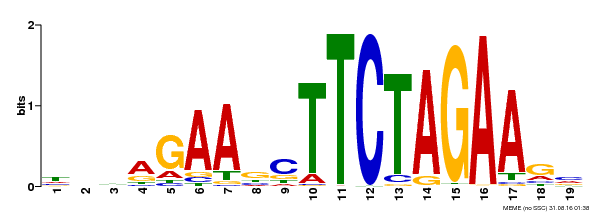
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010916598.1 | 0.0 | heat stress transcription factor A-4b | ||||
| Swissprot | Q94J16 | 1e-162 | HFA4B_ORYSJ; Heat stress transcription factor A-4b | ||||
| TrEMBL | A0A2H3ZHW9 | 0.0 | A0A2H3ZHW9_PHODC; heat stress transcription factor A-4b-like | ||||
| STRING | XP_008776668.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3296 | 37 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 1e-85 | heat shock transcription factor A4A | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



