 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010912147.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 595aa MW: 65781.2 Da PI: 8.0557 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.5 | 9.8e-13 | 448 | 494 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
XP_010912147.1 448 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAIAYITELQ 494
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 6.0E-50 | 58 | 244 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.85 | 444 | 493 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 5.1E-19 | 447 | 512 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 5.25E-14 | 447 | 498 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 2.7E-19 | 448 | 514 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.4E-10 | 448 | 494 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.0E-16 | 450 | 499 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS51671 | 9.476 | 530 | 595 | IPR002912 | ACT domain |
| CDD | cd04873 | 0.00701 | 543 | 587 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008152 | Biological Process | metabolic process | ||||
| GO:0016597 | Molecular Function | amino acid binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 595 aa Download sequence Send to blast |
MKTEMGTGVG VGVGLSVWSD EDRAMAVAVL GHQAVDYLFS SHVSSDGLLT AIGGDADLQN 60 KFQDLVDRPG SPWASAIFWQ ISRSKSGDLV LGWGDGHCRE LRDGEEDLRR SAAALDDGRQ 120 KMRKRVLQKL HVLSGGADDE NYALRLDRVT DVEMFFLASM YFSFPRGTGG PGKALASREH 180 VWVSDPSLRL PTLAADYCVR AFLARSAGFR TVVLVPFDTG VLELGSVKTV PESFEALQMI 240 RAVFSAGEKK DENGLVFRRV SASEEFPRIF GKDLNIGRRS QFNDRLSVAT KVEERPFEIQ 300 PNGGNHHRMV PVPVSSGPVN GLNWTQQQKF ANGMVVIGGG EVDPSSRAFV HHSNGVRDDP 360 RMNLFSTPKQ QQQQQQQQPP PPQPRQIDFS GGATSRASGA LAARLGAMES EHSDAEASCK 420 EERPGAMDER RPRKRGRKPA NGREEPLNHV EAERQRREKL NQRFYALRAV VPNISKMDKA 480 SLLGDAIAYI TELQKKLKEM EAERERWPES TSRLGQAPEI DVQAVHDEVI VRVSCPMDTH 540 PVSKVIQVFK ESQINVVDSK VSASNETVLH TFVVKSPGSE QLTRDKLVAA ISRER |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 9e-29 | 442 | 504 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 9e-29 | 442 | 504 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 9e-29 | 442 | 504 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 9e-29 | 442 | 504 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 9e-29 | 442 | 504 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 9e-29 | 442 | 504 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 9e-29 | 442 | 504 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 9e-29 | 442 | 504 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 429 | 437 | RRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
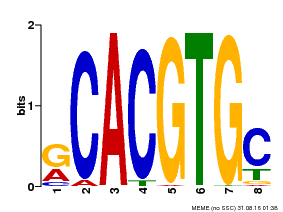
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010912147.1 | 0.0 | transcription factor bHLH13 | ||||
| Swissprot | A0A3Q7ELQ2 | 0.0 | MTB1_SOLLC; Transcription factor MTB1 | ||||
| TrEMBL | A0A2H3ZQG2 | 0.0 | A0A2H3ZQG2_PHODC; transcription factor bHLH13-like | ||||
| STRING | XP_008790159.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5174 | 38 | 60 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 1e-150 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



