 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010909816.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 437aa MW: 47200.8 Da PI: 7.042 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.2 | 1.3e-12 | 270 | 315 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr+++P+ +++Ka+ L Av YIk L
XP_010909816.1 270 NHVEAERQRREKLNHRFYALRSVVPNV-----SRMDKASLLADAVSYIKVL 315
799***********************6.....4***************977 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 9.9E-38 | 34 | 201 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| SuperFamily | SSF47459 | 2.49E-18 | 264 | 330 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.085 | 266 | 315 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 9.84E-15 | 269 | 320 | No hit | No description |
| Pfam | PF00010 | 3.7E-10 | 270 | 315 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 4.5E-18 | 270 | 333 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 6.0E-16 | 272 | 321 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005509 | Molecular Function | calcium ion binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 437 aa Download sequence Send to blast |
MEELISPSSS CSPSPPPPFL SIPTVQTAAV GPNLQQRLQY LLHARSEWWA YAIFWRASPD 60 HHLLSFGEGH FRGTRDVDRK PHGLRSRVGG IQALLSEVTG TTAGDIASDG DDVEWFYVVS 120 LTRSFAAQEP VLPARAYGSL APVWVAGAHA LQTCGCDRAR EAQLHGIETL VCVPLSGGVL 180 ELGSSDLIGE NWVLVQQAKA ILSAPDDAIV GPMVPAPPLV MNKVAGAGIT GLSSSVDSEH 240 SDSEGALMME QRRPKKRGRK PGTGRETPVN HVEAERQRRE KLNHRFYALR SVVPNVSRMD 300 KASLLADAVS YIKVLRAKVE ELEAEAKRIK KEVIGDQGVG GAATTSTTTT VSSGPLTMEL 360 EVKMLGPDAL IRVHSDNLNH PTAKLMGALR DLEVNVHHAS VSTVKEVMLQ DVVVRVPYAL 420 QGDDSLRAAL LSRLEKN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rqw_A | 2e-24 | 34 | 203 | 9 | 193 | Transcription factor MYC3 |
| 4rqw_B | 2e-24 | 34 | 203 | 9 | 193 | Transcription factor MYC3 |
| 4rs9_A | 2e-24 | 34 | 203 | 9 | 193 | Transcription factor MYC3 |
| 4yz6_A | 2e-24 | 34 | 203 | 9 | 193 | Transcription factor MYC3 |
| 5gnj_A | 2e-25 | 264 | 333 | 2 | 71 | Transcription factor MYC2 |
| 5gnj_B | 2e-25 | 264 | 333 | 2 | 71 | Transcription factor MYC2 |
| 5gnj_E | 2e-25 | 264 | 333 | 2 | 71 | Transcription factor MYC2 |
| 5gnj_F | 2e-25 | 264 | 333 | 2 | 71 | Transcription factor MYC2 |
| 5gnj_G | 2e-25 | 264 | 333 | 2 | 71 | Transcription factor MYC2 |
| 5gnj_I | 2e-25 | 264 | 333 | 2 | 71 | Transcription factor MYC2 |
| 5gnj_M | 2e-25 | 264 | 333 | 2 | 71 | Transcription factor MYC2 |
| 5gnj_N | 2e-25 | 264 | 333 | 2 | 71 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 251 | 259 | RRPKKRGRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00547 | DAP | Transfer from AT5G46830 | Download |
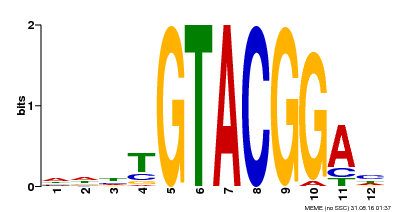
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010909816.1 | 0.0 | transcription factor MYC2 | ||||
| TrEMBL | A0A2H3X8N3 | 0.0 | A0A2H3X8N3_PHODC; transcription factor MYC2-like | ||||
| STRING | XP_008780029.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP893 | 33 | 130 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00870.1 | 7e-63 | bHLH family protein | ||||



