 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010558714.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 395aa MW: 44436 Da PI: 7.0591 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 107.8 | 5.7e-34 | 46 | 100 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kprl+Wtp+LHerFveav+qLGG++kAtPk+i+++m+++gLtl+h+kSHLQkYRl
XP_010558714.1 46 KPRLKWTPDLHERFVEAVNQLGGADKATPKAIMKVMGIPGLTLYHLKSHLQKYRL 100
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 10.777 | 43 | 103 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.1E-32 | 44 | 101 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.15E-17 | 45 | 101 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.9E-24 | 46 | 101 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.1E-10 | 48 | 99 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 5.3E-20 | 147 | 191 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 395 aa Download sequence Send to blast |
MYYHNQHQAK NTISTPRMPM PSERHPFLRG GNSPGDSGLI LSTDAKPRLK WTPDLHERFV 60 EAVNQLGGAD KATPKAIMKV MGIPGLTLYH LKSHLQKYRL SKNLNGQANS VSNKMGLMAM 120 AEEMSPDTSE IHTESLSIGP QTNKNLPISE ALQIQIEVQR RLHEQLERHL QLRIEAQGKY 180 LQAVLEKAQE TLGKQNLGTA GIEAAKVQLS ELASKVSAEY SNANFVEPKE LQSLHPQQMQ 240 MTYPPDFSLD SSLASREGTQ RSPKMLENRL GLRPYLSDST TEQKEIAEEP LFQRMELTWR 300 EGLRENKTYI SPMADDRETR LSSSERCPGN LSIGVGLHGH RGHGNADYAE ERIKKDEERI 360 QKGLDSTTEL DLNTHDGNYG TARPKQFDLN GFSWN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 5e-22 | 45 | 102 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 5e-22 | 45 | 102 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 5e-22 | 46 | 102 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 5e-22 | 46 | 102 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 5e-22 | 46 | 102 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 5e-22 | 46 | 102 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 5e-22 | 45 | 102 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 5e-22 | 45 | 102 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 5e-22 | 45 | 102 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 5e-22 | 45 | 102 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 5e-22 | 45 | 102 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 5e-22 | 45 | 102 | 1 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that may activate the transcription of specific genes involved in nitrogen uptake or assimilation (PubMed:15592750). Acts redundantly with MYR1 as a repressor of flowering and organ elongation under decreased light intensity (PubMed:21255164). Represses gibberellic acid (GA)-dependent responses and affects levels of bioactive GA (PubMed:21255164). {ECO:0000269|PubMed:21255164, ECO:0000305|PubMed:15592750}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00331 | DAP | Transfer from AT3G04030 | Download |
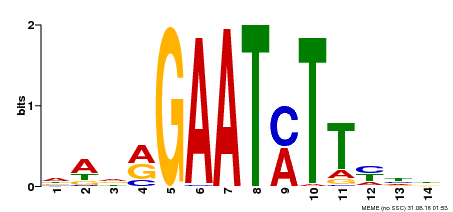
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010558714.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by nitrogen deficiency. {ECO:0000269|PubMed:15592750}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010558714.1 | 0.0 | PREDICTED: myb-related protein 2-like isoform X2 | ||||
| Swissprot | Q9SQQ9 | 0.0 | PHL9_ARATH; Myb-related protein 2 | ||||
| TrEMBL | D7L209 | 0.0 | D7L209_ARALL; Predicted protein | ||||
| STRING | XP_010558713.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G04030.3 | 0.0 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



