 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010545672.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Cleomaceae; Tarenaya
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 839aa MW: 92093.3 Da PI: 6.0823 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.7 | 2.1e-18 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
XP_010545672.1 17 KYVRYTPEQVEALERLYHDCPKPSSIRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 170.2 | 1.3e-53 | 161 | 369 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlm 94
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+++d+ W + ++++++l+v+ + g+++l
XP_010545672.1 161 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIIAISHGCTGVAARACGLVGLEPT-RVAEIVKDRSSWFRECRAVDVLNVLPTTngGTIELL 254
7899******************************************************.7777777777***************87667****** PP
EEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHH CS
START 95 vaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslv 185
+++l+a+++l+p Rdf+ +Ry+ l++g++v++++S+ ++q+ p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl++++++++lr+l+
XP_010545672.1 255 YMQLYAPTTLAPpRDFWLLRYTSVLEDGSLVVCERSLKNTQSGPNmppVQHFVRAEMLPSGYLIRPCEGGGSIIHIVDHMDLEASSVPEVLRPLY 349
******************************************99998899********************************************* PP
HHHHHHHHHHHHHHTXXXXX CS
START 186 ksglaegaktwvatlqrqce 205
+s + ++kt++a l+++++
XP_010545672.1 350 ESPKVLAQKTTMASLRQLKQ 369
***************99886 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.467 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-15 | 14 | 80 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.11E-16 | 16 | 79 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.32E-16 | 17 | 77 | No hit | No description |
| Pfam | PF00046 | 5.8E-16 | 18 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 3.5E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 3.56E-6 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 26.025 | 151 | 399 | IPR002913 | START domain |
| CDD | cd08875 | 1.06E-73 | 155 | 371 | No hit | No description |
| SuperFamily | SSF55961 | 2.33E-36 | 160 | 372 | No hit | No description |
| SMART | SM00234 | 1.3E-38 | 160 | 370 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 2.1E-22 | 160 | 365 | IPR023393 | START-like domain |
| Pfam | PF01852 | 5.6E-51 | 161 | 369 | IPR002913 | START domain |
| Pfam | PF08670 | 1.6E-51 | 697 | 838 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 839 aa Download sequence Send to blast |
MEMSYKDGKA GSLDNGKYVR YTPEQVEALE RLYHDCPKPS SIRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQSVNRKMT AMNKLLMEEN DRLQKQVSQL VYENSYFRQH 120 TQNTSLPAKD PSCESAVTSG QQQLASQNPP RDASPAGLLS IAEETLAEFL SKATGTAVEW 180 VQMPGMKPGP DSIGIIAISH GCTGVAARAC GLVGLEPTRV AEIVKDRSSW FRECRAVDVL 240 NVLPTTNGGT IELLYMQLYA PTTLAPPRDF WLLRYTSVLE DGSLVVCERS LKNTQSGPNM 300 PPVQHFVRAE MLPSGYLIRP CEGGGSIIHI VDHMDLEASS VPEVLRPLYE SPKVLAQKTT 360 MASLRQLKQI AQEVSQTSVS NGWSRRPAAL RALSQRLSRG FNEAVNGFTD EGWSVIGDGM 420 DDVTILVNSS PDKLMDLNLT FANNGFPHVS NAVLCAKASM LLQNVSPAIL LRFLREHRSE 480 WADNSIDAYS AASIKLGPCS LPGAWVGGCG GQVILPLAHT IEHEEFLEVI KLEGLGHSPE 540 DAIVPRDIFL LQLCSGIDES AVGTCGELIF APIDASFADD APLLPSGFRI IPLDLGKQEV 600 PSPNRTLDLA SALEVGSAGT KASTDGSGNS SCTRSVMTIA FEFAFESHMQ DHVVSMARQY 660 VRSIISTVQR VALALSPSHM SSQAGLLTSL GTPEAQTLAR WICHSYRCFM GVELLKSSDG 720 NESILKNLWN HSDAIMCCSL KALPVFTFAN QAGLDMLETT LVALQDISLE KIFDDNGRKT 780 LCSEFPQIMQ QGFACLRGGI CLSSMGRPVS YDKAVAWKVL NEEENAHCIC FVFINWSFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
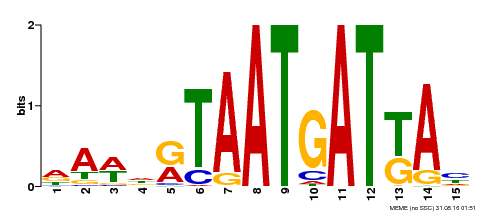
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010545672.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010545670.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15 isoform X1 | ||||
| Refseq | XP_010545671.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15 isoform X1 | ||||
| Refseq | XP_010545672.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein ATHB-15 isoform X1 | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | B3H4G8 | 0.0 | B3H4G8_ARATH; Homeobox-leucine zipper family protein / lipid-binding START domain-containing protein | ||||
| STRING | XP_010545669.1 | 0.0 | (Tarenaya hassleriana) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.2 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



