 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010112795.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Moraceae; Morus
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 349aa MW: 38875.3 Da PI: 9.497 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 92.9 | 2.3e-29 | 36 | 93 | 2 | 60 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS-- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhek 60
+DgynWrKYGqK vkg+e+ rsYYrCt+++C+vkk++e+s+ d ++v+i+Y g+H+h+k
XP_010112795.1 36 QDGYNWRKYGQKLVKGNEYVRSYYRCTHPNCQVKKQLECSH-DRQIVDIVYFGHHDHPK 93
8***************************************9.***************85 PP
| |||||||
| 2 | WRKY | 103 | 1.7e-32 | 210 | 268 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
++Dg++WrKYGqK vkgs++prsYYrC+ +gCpvkk+ver+++d kvv++tYegeHnh
XP_010112795.1 210 VNDGHRWRKYGQKIVKGSPHPRSYYRCSNSGCPVKKHVERASHDAKVVITTYEGEHNHG 268
58********************************************************5 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 1.8E-25 | 29 | 93 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 20.748 | 30 | 94 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 2.48E-23 | 31 | 93 | IPR003657 | WRKY domain |
| SMART | SM00774 | 6.5E-31 | 35 | 93 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 2.1E-22 | 36 | 92 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 5.8E-36 | 196 | 270 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.7E-29 | 202 | 270 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 35.026 | 205 | 270 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.3E-38 | 210 | 269 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 6.3E-27 | 211 | 267 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009863 | Biological Process | salicylic acid mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 349 aa Download sequence Send to blast |
MPVPEKASQS PETGVLALKS VQEGSTPSII REKVAQDGYN WRKYGQKLVK GNEYVRSYYR 60 CTHPNCQVKK QLECSHDRQI VDIVYFGHHD HPKRHLNNNI PLAVGFVVST AEQRRKEPPL 120 TTTEDVSKNE RSLALTKTER VDIPPVTNIA ANDSPKGITQ SNKIKEVKSE DDPASKRQKK 180 DEHDHKTTSV DKPINEARLV VQTRSEVDIV NDGHRWRKYG QKIVKGSPHP RSYYRCSNSG 240 CPVKKHVERA SHDAKVVITT YEGEHNHGMP PTRIVTHNVP GSGTSPTAND GDTGTKSDAR 300 ALLEDKSESK EHANRELGTK SAVNETTGSN IESCPSPDSK TVKQENGKK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ayd_A | 2e-36 | 34 | 271 | 1 | 75 | WRKY transcription factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Binds to a 5'-CGTTGACCGAG-3' consensus core sequence which contains a W box, a frequently occurring elicitor-responsive cis-acting element. {ECO:0000269|PubMed:8972846}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00094 | SELEX | Transfer from AT2G04880 | Download |
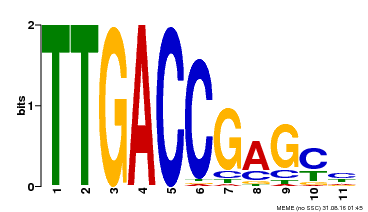
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010112795.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salicylic acid (SA). {ECO:0000269|PubMed:17264121}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024032086.1 | 0.0 | WRKY transcription factor 1 isoform X1 | ||||
| Refseq | XP_024032087.1 | 0.0 | WRKY transcription factor 1 isoform X1 | ||||
| Refseq | XP_024032088.1 | 0.0 | WRKY transcription factor 1 isoform X1 | ||||
| Swissprot | Q9SI37 | 4e-88 | WRKY1_ARATH; WRKY transcription factor 1 | ||||
| TrEMBL | W9SZV2 | 0.0 | W9SZV2_9ROSA; WRKY transcription factor 1 | ||||
| STRING | XP_010112795.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF9358 | 33 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G04880.2 | 2e-92 | zinc-dependent activator protein-1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 21402940 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



