 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010105257.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Moraceae; Morus
|
||||||||
| Family | BBR-BPC | ||||||||
| Protein Properties | Length: 337aa MW: 37487.4 Da PI: 10.0207 | ||||||||
| Description | BBR-BPC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GAGA_bind | 397.5 | 1.9e-121 | 1 | 337 | 1 | 301 |
GAGA_bind 1 mdddgsre..rnkg.yyepa..aslkenl..glqlmssiaerdakirernlalsekkaavaerdmaflqrdkalaernkalverdnkllalllve 88
mdd g+re r+kg +y+ a + + +++ +q+m+++aerda+i+ernlal+ekkaa+aerdmaflqrd+a+aern+a++erdn++++l+++e
XP_010105257.1 1 MDDGGHREngRHKGdQYKTAqgQWMMQHQpsMKQIMALMAERDAAIQERNLALAEKKAALAERDMAFLQRDAAIAERNNAIMERDNAVATLQYRE 95
9*****999*********555544444442246************************************************************** PP
GAGA_bind 89 nsla....salpvgvqvlsgtksidslqqlse..pqledsavelreeeklealpieeaaeeakekkkkkkrqrakkpkekkakkkkkksekskkk 177
nsl+ s++p+g+q+++g+k++++ q++ + +++++ ++ +re++++++lpi+ a++a ++++ k+ ++a k +++kk +k+ +k
XP_010105257.1 96 NSLSngnmSSCPPGCQISRGVKHMHHPQTHAHhmQHVNEGSYATREMNTTDSLPISPDASDAAKSRRGKRMKEA------KPVSPNKKASKPPRK 184
**9999999*******************955557889999***************9988777766655543333......333333333333333 PP
GAGA_bind 178 vkkesader.............................skaekksidlvlngvslDestlPvPvCsCtGalrqCYkWGnGGWqSaCCtttiSvyP 243
k+es+d + sk+++k +dl+ln+vs+Dest+P+PvC+CtG+ rqCYkWGnGGWqS+CCttt+S+yP
XP_010105257.1 185 LKRESEDLNkmafgkaqewksgqgmvggsddfnkqlvvSKSDWKGQDLGLNQVSYDESTMPAPVCTCTGVIRQCYKWGNGGWQSSCCTTTMSMYP 279
333333322233444456889************************************************************************** PP
GAGA_bind 244 LPvstkrrgaRiagrKmSqgafkklLekLaaeGydlsnpvDLkdhWAkHGtnkfvtir 301
LP+++++r+aR++grKmS++af+klL++LaaeG+dlsnpvDLkdhWAkHGtn+++ti+
XP_010105257.1 280 LPAVPNKRHARVGGRKMSGSAFNKLLSRLAAEGHDLSNPVDLKDHWAKHGTNRYITIK 337
*********************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01226 | 1.6E-181 | 1 | 337 | IPR010409 | GAGA-binding transcriptional activator |
| Pfam | PF06217 | 1.1E-104 | 1 | 337 | IPR010409 | GAGA-binding transcriptional activator |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 337 aa Download sequence Send to blast |
MDDGGHRENG RHKGDQYKTA QGQWMMQHQP SMKQIMALMA ERDAAIQERN LALAEKKAAL 60 AERDMAFLQR DAAIAERNNA IMERDNAVAT LQYRENSLSN GNMSSCPPGC QISRGVKHMH 120 HPQTHAHHMQ HVNEGSYATR EMNTTDSLPI SPDASDAAKS RRGKRMKEAK PVSPNKKASK 180 PPRKLKRESE DLNKMAFGKA QEWKSGQGMV GGSDDFNKQL VVSKSDWKGQ DLGLNQVSYD 240 ESTMPAPVCT CTGVIRQCYK WGNGGWQSSC CTTTMSMYPL PAVPNKRHAR VGGRKMSGSA 300 FNKLLSRLAA EGHDLSNPVD LKDHWAKHGT NRYITIK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds to GA-rich elements (GAGA-repeats) present in regulatory sequences of genes involved in developmental processes. {ECO:0000269|PubMed:14731261}. | |||||
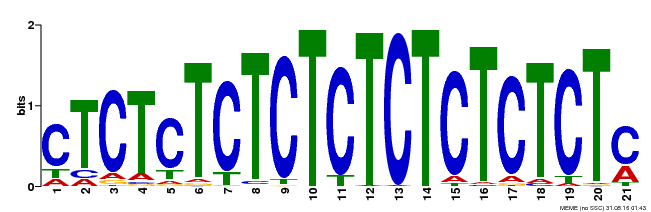
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00540 | DAP | Transfer from AT5G42520 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010105257.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010105257.1 | 0.0 | protein BASIC PENTACYSTEINE6 | ||||
| Swissprot | Q8L999 | 1e-160 | BPC6_ARATH; Protein BASIC PENTACYSTEINE6 | ||||
| TrEMBL | W9RR46 | 0.0 | W9RR46_9ROSA; Uncharacterized protein | ||||
| STRING | XP_010105257.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF6905 | 34 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42520.1 | 1e-154 | basic pentacysteine 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 21402340 |



