 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010101942.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Moraceae; Morus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1508aa MW: 169200 Da PI: 8.6449 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.4 | 0.00022 | 1393 | 1418 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y C C++sFs+k +L+ H ++ +
XP_010101942.1 1393 YMCDieGCTMSFSTKQELVLHKKNiC 1418
899999***************99866 PP
| |||||||
| 2 | zf-C2H2 | 13.1 | 0.00028 | 1418 | 1440 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk F ++ +L++H r+H
XP_010101942.1 1418 CPvkGCGKKFFSHKYLVQHRRVH 1440
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 11.8 | 0.00074 | 1476 | 1502 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
XP_010101942.1 1476 YVCAepGCGQTFRFVSDFSRHKRKtgH 1502
899999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.5E-14 | 24 | 65 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.55 | 25 | 66 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 7.8E-14 | 26 | 59 | IPR003349 | JmjN domain |
| PROSITE profile | PS51184 | 33.514 | 186 | 365 | IPR003347 | JmjC domain |
| SMART | SM00558 | 2.6E-50 | 196 | 365 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 3.3E-26 | 210 | 379 | No hit | No description |
| Pfam | PF02373 | 2.0E-36 | 229 | 348 | IPR003347 | JmjC domain |
| SMART | SM00355 | 10 | 1393 | 1415 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 0.0045 | 1416 | 1440 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 7.21E-6 | 1416 | 1452 | No hit | No description |
| PROSITE profile | PS50157 | 12.445 | 1416 | 1445 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-5 | 1417 | 1444 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1418 | 1440 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-7 | 1445 | 1470 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0017 | 1446 | 1470 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.637 | 1446 | 1475 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1448 | 1470 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.68E-9 | 1456 | 1500 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.3E-9 | 1471 | 1499 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.011 | 1476 | 1507 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.62 | 1476 | 1502 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1478 | 1502 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0032259 | Biological Process | methylation | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0040010 | Biological Process | positive regulation of growth rate | ||||
| GO:0045815 | Biological Process | positive regulation of gene expression, epigenetic | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0008168 | Molecular Function | methyltransferase activity | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0071558 | Molecular Function | histone demethylase activity (H3-K27 specific) | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1508 aa Download sequence Send to blast |
MAASGLTSEQ ASPEVFSWLK TLPQAPEYHP TLAEFQDPIS YIFKIEKEAS EYGICKIVPP 60 VPPSAKKTVI ANLNKSLAAR NGGFDASNPK NPPTFTTRQQ QIGFCPRKPR PVQRPVWQSG 120 ENYTFQQFEA KAKGFERSFF KRCAKKGALS PLEIETLYWK ATVDKPFSVE YANDMPGSAF 180 VPVSAKRSRE AGESATLGET AWNMRAVSRA KGSLLRFMKE EIPGVTSPMV YVAMMFSWFA 240 WHVEDHDLHS LNYLHMGAGK TWYGVPREAA VAFEEVVRVH GYGGEINPLV TFSILGEKTT 300 VMSPEVFVRA GVPCCRLVQN PGEFVVTFPR AYHTGFSHGF NCGEAANIAT PEWLRVAKDA 360 AIRRASINYP PMVSHFQLLY DLALALCSRI PESVGAEPRS SRLKDKKKGE GETVVKELFV 420 QNVLQNNDLL HVLGNGSPVV LLPRSSSDIS VCSKLRVGSH LRLNSSSPLA SCNSREEMKS 480 SRSLISDDLM IDRKQEVDQV KDFYSVKGKL ASLCDRSWVP SLRGNKITCA SNSKTSNMNV 540 EGESTVDNDG LSDQRLFSCV TCGILSFACV AIIQPREPAA RYLMSADCSF FNDWVVNAGV 600 ASNVFPVSNR YQTASKENTY TGWTDNSEPL ALCENPGQSV NFQAQMADQK NEIVSNTETQ 660 KAPSALGLLA LNYGNSSDSE EDQVQEDVSV DGNETNVSNC SLESKYRCES SSPSLRNCQG 720 DTVHGRSLVE LDSGDDFASQ NADSYMENGH NKDNTKYDSH QNFDCPVSFR TNNAAPAQSN 780 GLVPKFGDGM KASRTCSPDT YDAEATRFCK AIAPTKNENM PFVPICDEDS CRMHVFCLEH 840 AVEVEQQLRQ VGCVDIVLLC HPDYPKIETE AKAMAEELGI SHLWNDIEFR DATKDDENMI 900 QATLDSEEAI PKNGDWAVKL GINLFYSANL SRSPLYSKQM PYNSVIYDAF GRSSPASSSA 960 RSDGFERRPA KQKKVVAGKW CGKVWMSSQV HPFLAKKDPE EEEQERSFHT WATPDEKVER 1020 KYDGTRKSSN TMIAKKYVRK RKMTVESSST KKAKRVKRED AVSDNSMDDS HEHHRRSLRS 1080 KQAVSIGGGS AKKAKHTEIE GAASDDSLHD NSHRQHRRTF KSKQATYVES DGIVSDDSLE 1140 VDFRYQHKKI LRSKPSKHAG REDVVSDDSL DSDSHQLRGR VCRIKQAKHT EEEDVVSDDS 1200 LDSDSQLHRS IPRSKQAKYN EREDSSSDYF HRNNLQKLHR RISKSKPAKS IGREDEDLDE 1260 PLEDNARKSD ERILRSKRTK SALQQKMKQE TPHHVKQSTA RPVKQENRKL KQQTPRLRNS 1320 QCEQNILGSC AEEELEGGPS TRLRKRNPKP QKLTGAKRKE QQQPSRKKVK NAVVVKAQAG 1380 HNDAKSKDEE GEYMCDIEGC TMSFSTKQEL VLHKKNICPV KGCGKKFFSH KYLVQHRRVH 1440 MDDRPLRCPW KGCKMTFKWA WARTEHIRVH TGARPYVCAE PGCGQTFRFV SDFSRHKRKT 1500 GHSVKKAR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 1e-77 | 18 | 386 | 8 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 1e-77 | 18 | 386 | 8 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1035 | 1041 | KYVRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
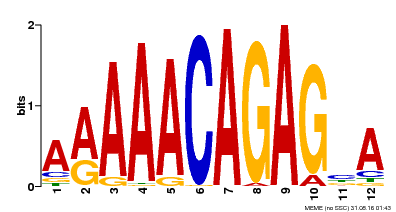
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010101942.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010101942.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | W9S5V7 | 0.0 | W9S5V7_9ROSA; Lysine-specific demethylase REF6 | ||||
| STRING | XP_010101942.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 21391565 |



