 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010099677.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Moraceae; Morus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 475aa MW: 52437.1 Da PI: 5.7736 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53.3 | 6.1e-17 | 60 | 104 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE++++++a k++G g W+ I +++g ++t+ q++s+ qk+
XP_010099677.1 60 REKWTEEEHQKFLEALKLYGRG-WRQIEEHVG-TKTAVQIRSHAQKF 104
789*******************.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.69E-16 | 54 | 107 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.622 | 55 | 109 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-9 | 57 | 107 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.7E-16 | 58 | 107 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.8E-13 | 59 | 107 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 9.4E-15 | 60 | 103 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.47E-11 | 62 | 105 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 475 aa Download sequence Send to blast |
MDGCDQNTYL AMQQSEGMEL KTSVVNGSKG DSKCESDNQM NEIYDTAPKV RKPYTITKQR 60 EKWTEEEHQK FLEALKLYGR GWRQIEEHVG TKTAVQIRSH AQKFFSKVVR GSTGTVESSI 120 KRIEIPPPRP KRKPVHPYPR KSAESFNGIL VSNQPERSPS PNLSAADKGT KSPTSVLSCQ 180 GSDALGSVAS GQHNRSPSSA SCTTDMNSIS QSPTEIENEY MIGNSSDEER RSLPAKHLSS 240 SLTLENLLSK KFELGAKDAS YTKEEEEEEE EEEEEAEVAA SASFKLFGRT VLLSGFQKES 300 PPGAKNSQTS KTDEGLVLTL SSDQLDTNLS LGGAVSNCNQ LEQPKESCNT ETANSPMLWW 360 TLYQFPFHYL AACNQTLDQT PRNSCSEHRE EDKDIPKERS STGSNDGSAS GVETAGDKNL 420 EAVDSFPQKP CTKVSVEPSN SRKGFVPYKR CLAERDNASA AVVPNVRERR KIRVC |
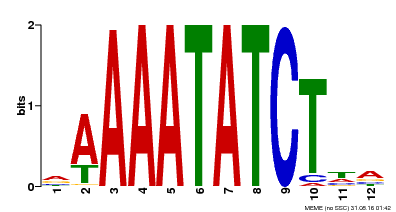
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_010099677.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KM001671 | 2e-60 | KM001671.1 Pyrus x bretschneideri transcription factor MYBR (MYBR) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024023521.1 | 0.0 | protein REVEILLE 7 isoform X1 | ||||
| TrEMBL | W9RAU0 | 0.0 | W9RAU0_9ROSA; Myb-like protein G | ||||
| STRING | XP_010099677.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7313 | 32 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.2 | 2e-46 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 21407819 |



