 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009757612.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Nicotianoideae; Nicotianeae; Nicotiana
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 483aa MW: 53580.7 Da PI: 6.791 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 54.1 | 2.8e-17 | 303 | 349 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn ErrRRdriN+++ L+ellP++ +K +Ka++L +A+eY+ksLq
XP_009757612.1 303 VHNLSERRRRDRINEKMKALQELLPHS-----TKTDKASMLDEAIEYLKSLQ 349
6*************************9.....9******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 3.53E-20 | 297 | 364 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.476 | 299 | 348 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 4.96E-18 | 302 | 353 | No hit | No description |
| Pfam | PF00010 | 1.2E-14 | 303 | 349 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 4.2E-20 | 303 | 357 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.0E-18 | 305 | 354 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0009693 | Biological Process | ethylene biosynthetic process | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 483 aa Download sequence Send to blast |
MNPCLPEWNI EAELPVHQKK PMGFDHELVE LLWRNGEVVL HSQTHKKQQG YDPNECRQFN 60 KHDQQTIRDI VSCGNHTSLI QDDETISWLN CPIDESFEKE FCSPFLSEIS TNPIEADKSI 120 RQSEDNKAFK FDPLEINHVF PHSHHSSFDP NPMPPPRFHN SVSAQKSVKN NLKEGSVMTV 180 GSSHCGSNQV AIDADTSRFS SSANIGLSAA MITDYTGKVS PQSDTMDRDT FEPANTSSSG 240 GSGSSYARTC NLSAATNSQS HKRKSRDGEE PECQSKAGEL ESAGGNKSAQ KSGTARRSRA 300 AEVHNLSERR RRDRINEKMK ALQELLPHST KTDKASMLDE AIEYLKSLQM QLQMMWMGSG 360 MASMMFPGVQ HYMSRIGMGM GSPTVPSIHN AMHLARLPLV DSTIPMNTAA PNQAAICQNS 420 MLNSVNYQRH LQNPNFQDQY ASYMGFHPLQ GASQQMNIFG LGSQTAQQSQ QLPHPTNSNA 480 PNT |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 307 | 312 | ERRRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00082 | ChIP-seq | Transfer from AT3G59060 | Download |
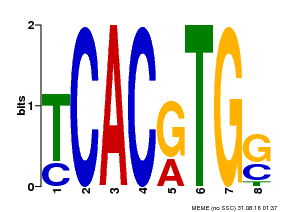
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT013582 | 0.0 | BT013582.1 Lycopersicon esculentum clone 132330F, mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009757612.1 | 0.0 | PREDICTED: transcription factor PIF4 | ||||
| Refseq | XP_009757613.1 | 0.0 | PREDICTED: transcription factor PIF4 | ||||
| Refseq | XP_016471316.1 | 0.0 | PREDICTED: transcription factor PIF4-like | ||||
| Refseq | XP_016471375.1 | 0.0 | PREDICTED: transcription factor PIF4-like | ||||
| TrEMBL | A0A1S4A419 | 0.0 | A0A1S4A419_TOBAC; transcription factor PIF4-like | ||||
| TrEMBL | A0A1U7UTT3 | 0.0 | A0A1U7UTT3_NICSY; transcription factor PIF4 | ||||
| STRING | XP_009757612.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA10465 | 22 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43010.2 | 7e-50 | phytochrome interacting factor 4 | ||||



