 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009148369.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 268aa MW: 29305.7 Da PI: 5.9754 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 33.3 | 8.8e-11 | 152 | 199 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L++l+P + +k Ka +L + ++YI+sLq
XP_009148369.1 152 SHSLAERARREKISERMKILQDLVPGC----NKVIGKALVLDEIINYIQSLQ 199
8*************************9....677*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 8.54E-11 | 146 | 203 | No hit | No description |
| SuperFamily | SSF47459 | 5.89E-18 | 146 | 217 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.828 | 148 | 198 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 4.3E-18 | 148 | 216 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.8E-8 | 152 | 199 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.0E-10 | 154 | 204 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048446 | Biological Process | petal morphogenesis | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 268 aa Download sequence Send to blast |
MDPGGMMNEA GAYNLAEIWP YPTINGGAAV NTSMRSGVSF GGPNLVHQFA DSNLISNTNC 60 NDPARMSHAL SQAVVAGIAG ARKRREAGAE DNNVVSGSDG HDANDGDSKK QKTTGCEDEV 120 EVGTLDQKKQ QLPEPPKDYI HVRARRGQAT DSHSLAERAR REKISERMKI LQDLVPGCNK 180 VIGKALVLDE IINYIQSLQR QVEFLSMKLE AVNSRMTPGI EVFPPKEFDQ QTFENQGMQF 240 GAQGTREYSR GASPEWLHMQ VGGGFERS |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bra.19016 | 0.0 | bud| flower | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Isoform 1 is specifically expressed in flowers, mostly in petals, inflorescence and flower buds. Isoform 2 is expressed ubiquitously (leaves, flowers and stems). {ECO:0000269|PubMed:16902407}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the control of petal size, by interfering with postmitotic cell expansion to limit final petal cell size. {ECO:0000269|PubMed:16902407}. | |||||
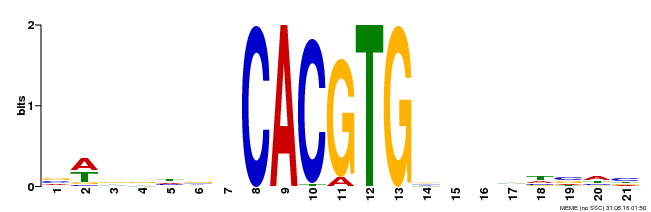
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00207 | DAP | Transfer from AT1G59640 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009148369.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Isoform 1 is up-regulated by PI/AP3, SEP2, SEP3 and AP1, but repressed by AG. {ECO:0000269|PubMed:16902407}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY086373 | 1e-109 | AY086373.1 Arabidopsis thaliana clone 24644 mRNA, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009148369.1 | 0.0 | PREDICTED: transcription factor BPE | ||||
| Swissprot | Q0JXE7 | 2e-96 | BPE_ARATH; Transcription factor BPE | ||||
| TrEMBL | M4DTU2 | 0.0 | M4DTU2_BRARP; Uncharacterized protein | ||||
| STRING | Bra019935.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1855 | 28 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G59640.1 | 1e-118 | BIG PETAL P | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



