 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009143403.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 337aa MW: 38652.9 Da PI: 8.6384 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.3 | 6.1e-32 | 57 | 110 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH rFv+ave+LGG+e+AtPk ++++m++kgL+++hvkSHLQ+YR+
XP_009143403.1 57 PRLRWTPDLHLRFVRAVERLGGQERATPKLVRQMMNIKGLSIAHVKSHLQMYRS 110
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 4.5E-29 | 53 | 111 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 12.273 | 53 | 113 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.79E-14 | 55 | 111 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-22 | 57 | 112 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.2E-9 | 58 | 109 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 337 aa Download sequence Send to blast |
MIPNMEGSVK TTDREEEEEE EEEEGEGEES KVSSNTTVEA EVGKKTKVRP YVRSKVPRLR 60 WTPDLHLRFV RAVERLGGQE RATPKLVRQM MNIKGLSIAH VKSHLQMYRS KKMDDQGQAI 120 ADNRHFIESS TDRNIYKLSQ LPMFRGYNTN HSHDSPFRYG SRFSNASLWN SSSHETNRSL 180 IDRTGLIRGS SVNNNIHGSE YWTNNRSFQN TYSSSVSNHL PKLRHNHQER NNLATFNSIQ 240 GHSRTFEKFE TGIEERTNHV YCTKTTGKRN ASTSLDLDLS LKLRVPEETT LEETETATTD 300 QTLSLSLCSW KKSRVIKTDE EDRTVKIGQA STLDLTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 9e-17 | 58 | 112 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 9e-17 | 58 | 112 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 9e-17 | 58 | 112 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 9e-17 | 58 | 112 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
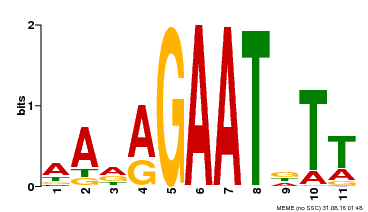
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009143403.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009143403.1 | 0.0 | PREDICTED: putative Myb family transcription factor At1g14600 | ||||
| TrEMBL | A0A3P5Z7B1 | 0.0 | A0A3P5Z7B1_BRACM; Uncharacterized protein | ||||
| TrEMBL | M4CLN7 | 0.0 | M4CLN7_BRARP; Uncharacterized protein | ||||
| STRING | Bra005124.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM363 | 28 | 184 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 1e-168 | G2-like family protein | ||||



