 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_009104878.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassiceae; Brassica
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 379aa MW: 43143.3 Da PI: 7.3961 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11.1 | 0.0013 | 21 | 43 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
y C++Cg s s k++ ++Hi +H
XP_009104878.1 21 YLCQYCGISRSKKYLITSHIESH 43
56*******************96 PP
| |||||||
| 2 | zf-C2H2 | 20.3 | 1.4e-06 | 68 | 89 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg F+ + +Lk+H+++H
XP_009104878.1 68 TCNECGAEFKKPAHLKQHMQSH 89
7*******************99 PP
| |||||||
| 3 | zf-C2H2 | 16.4 | 2.6e-05 | 95 | 119 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ C dC+ s++rk++L+rH+ tH
XP_009104878.1 95 FACYvdDCSSSYRRKDHLNRHLLTH 119
678888*****************99 PP
| |||||||
| 4 | zf-C2H2 | 20 | 1.8e-06 | 182 | 205 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+C+ Cgk F+ +s+L++H +H
XP_009104878.1 182 VCKevGCGKAFKYPSQLQKHEDSH 205
79999***************9888 PP
| |||||||
| 5 | zf-C2H2 | 20.3 | 1.5e-06 | 272 | 297 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
+kC+ C+ +Fs +snL++H++ H
XP_009104878.1 272 FKCEveGCSSTFSKPSNLQKHLKAvH 297
89********************9877 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 26 | 21 | 43 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.464 | 67 | 94 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0025 | 67 | 89 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 7.0E-5 | 68 | 85 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 69 | 89 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.18E-11 | 76 | 128 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 5.6E-13 | 86 | 121 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 9.972 | 95 | 124 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.21 | 95 | 119 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 97 | 119 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.9E-5 | 122 | 150 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 6.0E-4 | 122 | 152 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 1.1 | 124 | 149 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.598 | 124 | 154 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 126 | 149 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 8.26E-5 | 177 | 206 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 9.4E-6 | 179 | 205 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0055 | 181 | 205 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.471 | 181 | 210 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 183 | 205 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 2.2 | 213 | 238 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 215 | 238 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 25 | 241 | 262 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 1.32E-8 | 252 | 307 | No hit | No description |
| PROSITE profile | PS50157 | 12.84 | 272 | 302 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3.6E-4 | 272 | 297 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 7.9E-8 | 272 | 297 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 274 | 297 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 8.3E-5 | 298 | 324 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 4.9 | 303 | 329 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 305 | 329 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 379 aa Download sequence Send to blast |
MSEEAANVDV NPTKTKDIRN YLCQYCGISR SKKYLITSHI ESHHQMEVEE ERNEEACEDE 60 EEISGKQTCN ECGAEFKKPA HLKQHMQSHS LERPFACYVD DCSSSYRRKD HLNRHLLTHK 120 GKLFKCPVEN CKSEFSIHGN ISKHLKKFHN NSDSNKDDTS NGDSVNGDSQ PSGCSSGQKK 180 LVCKEVGCGK AFKYPSQLQK HEDSHVKLDS VEAFCSEIGC MKYFTNEECL KAHIRSCHQH 240 VNCEICGSKH LKKNIKRHLR THDEDSSSPG EFKCEVEGCS STFSKPSNLQ KHLKAVHEDI 300 RPFVCGFPGC GMRFAYKHVR NNHENSGSHV YTCGDFVEAD EDFTSRPRGG LKRKQVTAEM 360 LVRKRVMPPQ FDSEEQETC |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wjq_D | 2e-17 | 19 | 332 | 7 | 284 | Zinc finger protein 568 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Bra.9255 | 0.0 | leaf | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Both isoforms 1 and 2 transcripts are expressed in seedlings and both TFIIIA and TFIIIA-C are present at similar ratios in 10-day-old seedlings (at protein level). In correlation with the reorganization of 5S rDNA chromatin to a mature state, full-length isoform 1 protein TFIIIA with transcriptional activity accumulates and permits de novo transcription of 5S rRNA. Isoforms 1 and 2 transcripts accumulate in various tissues of the reproductive phase, including flowers and siliques, but only isoform 2 is present in mature seeds. In flowers, both TFIIIA and TFIIIA-C are present at similar ratios (at protein level). Very low amounts of TFIIIA are found in extracts from fresh siliques compared to TFIIIA-C. The amount of full-length TFIIIA protein progressively increases from fresh siliques to seeds concomitant with lower proportions of the shorter TFIIIA-C protein (at protein level) thus leading to 5S rRNA accumulation in the seed. {ECO:0000269|PubMed:22353599}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in seedlings, flowers, siliques and seeds. {ECO:0000269|PubMed:22353599}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
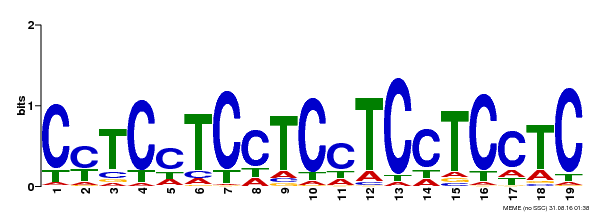
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_009104878.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY054225 | 0.0 | AY054225.1 Arabidopsis thaliana At1g72050/F28P5_6 mRNA, complete cds. | |||
| GenBank | AY066042 | 0.0 | AY066042.1 Arabidopsis thaliana At1g72050/F28P5_6 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009104878.1 | 0.0 | PREDICTED: transcription factor IIIA-like | ||||
| Swissprot | Q84MZ4 | 0.0 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | M4CI66 | 0.0 | M4CI66_BRARP; Uncharacterized protein | ||||
| STRING | Bra003899.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4714 | 28 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 0.0 | transcription factor IIIA | ||||



