 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_008237378.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1086aa MW: 121454 Da PI: 5.7628 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 185.3 | 6.4e-58 | 21 | 136 | 4 | 118 |
CG-1 4 e.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrr 97
e ++rwl++ ei++iL nf+k+++++e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fqrr
XP_008237378.1 21 EaQHRWLRPAEICEILSNFQKFHISSEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQRR 115
449******************************************************************************************** PP
CG-1 98 cywlLeeelekivlvhylevk 118
+yw+Le++l +iv+vhylevk
XP_008237378.1 116 SYWMLEQDLMHIVFVHYLEVK 136
******************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 85.995 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.4E-85 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.3E-51 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 4.5E-5 | 487 | 589 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 9.1E-17 | 503 | 588 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 17.926 | 669 | 768 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 2.1E-6 | 685 | 762 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 5.91E-17 | 687 | 797 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.97E-14 | 687 | 795 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 1.8E-16 | 691 | 798 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 3400 | 703 | 732 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0015 | 736 | 765 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 10.286 | 736 | 768 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 7.9 | 909 | 931 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 1.38E-7 | 911 | 960 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 7.95 | 911 | 939 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0023 | 912 | 930 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0071 | 932 | 954 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.212 | 933 | 957 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.3E-4 | 935 | 954 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1086 aa Download sequence Send to blast |
MAERGSYSQG PRLDFQQLLG EAQHRWLRPA EICEILSNFQ KFHISSEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHEK LKVGSVDVLH CYYAHGEENE NFQRRSYWML 120 EQDLMHIVFV HYLEVKGNRA NAGGIREIDE VTPDLQKGSF WTSSPSNSNC RTPSGNTDYT 180 SPSSNLTSCE DADSGDSRQA SSFQSSFDSQ QMGNGPLTDK ADINLSLHPH LNNHDGQSSF 240 HGGNYRPYFE KDQQCYSDDS TCGIDSQKTL GVGSWEEILE QCTTGLHPVT SHGSKSSIQI 300 ASAGIPPEQQ NIASTEFLAG NSTLKEEFGS PLPFRTNWQI PLEENALQLP KWPLDQSMNL 360 QLPSNLDTRL FEQGTVDVNL RNAPELVTTH PNQRNDQLVQ NNFQAQLTNA ESQCLIISSS 420 EPDIPKDGNI NYAFTLRQQL LDQEEGLKKV DSFSRWVSKE LGEVDDLQMQ SSSGISWSTD 480 ECGNVADDSS LSPSISQDQL FSIVDFSPKW AYTDSEIEVL VIGTFLVSQK EVTKYNWSCM 540 FGEVEVPAQV LANGVLFCFA PPHSAGQVPF YVTCSNRLAC SEVREFEYQV GSTKDLDITN 600 ICNATTNDIH LHLRLESLLS LRSVSPSGQL VEGVKEKQNL ISKIISLKEE EECLPLVEPT 660 AVNDLLQHEG MEHLIKLMKE KLYSWLLHKA IEDGKGPSVL DSEGQGVIHL AAALGYDWAI 720 KPIVTAGVSI NFRDVNGWTA LHWAAFYGRE QTVAILISLG AAPGALTDPS PEFPLGRAPA 780 DLASVNRHKG ISGFLAESSL TSYLDSLTMN DAKEGGAAEI SRIRAVKTFS ERIATPGSYS 840 DMPDALSLKD SLTAVTNATQ AADRIHQMFR MQSFDRRQLT EYDTDEFGMP DERAISLIAS 900 KSHKVGPVNG HTAAIQIQKK FRGWKKRKEF LIIRQRIVKI QAHVRGHQVR KQYKAITWSV 960 GILEKVILRW RRKGTGLRGF RPDAVAKAPN LQSVPSNDDD YDFLKKGRKQ TEERLQKALT 1020 RVKSMVQYPE GRAQYRRLLN VVEGFRETKV SDMATDGSEL KVEGGDDLID IDKLLDDDTF 1080 MSIAFD |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
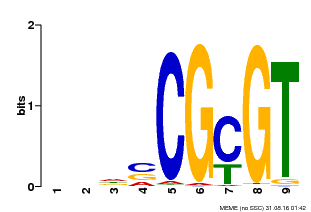
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_008237378.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008237378.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A251MWZ5 | 0.0 | A0A251MWZ5_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008237378.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7406 | 26 | 42 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



