 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_008236260.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 523aa MW: 57779.4 Da PI: 6.5106 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 45.6 | 1.8e-14 | 235 | 294 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + g ++ ++++lg ++ +++Aa+a++ a++k++g
XP_008236260.1 235 SIYRGVTRHRWTGRYEAHLWDnSCRrEGqaRKgRQVYLGGYDKEDKAARAYDLAALKYWG 294
57*******************666664478446**********99*************98 PP
| |||||||
| 2 | AP2 | 49.8 | 8.3e-16 | 337 | 388 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
XP_008236260.1 337 SIYRGVTRHHQQGRWQARIGRVAG---NKDLYLGTFATEEEAAEAYDIAAIKFRG 388
57****************988532...5************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 2.94E-17 | 235 | 304 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 1.30E-22 | 235 | 304 | No hit | No description |
| Pfam | PF00847 | 2.2E-11 | 235 | 294 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 9.0E-16 | 236 | 303 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 3.2E-29 | 236 | 308 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.137 | 236 | 302 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.4E-6 | 237 | 248 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 3.5E-10 | 337 | 388 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.71E-11 | 337 | 396 | No hit | No description |
| SuperFamily | SSF54171 | 9.15E-18 | 337 | 397 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 4.7E-18 | 338 | 396 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.111 | 338 | 396 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 7.9E-31 | 338 | 402 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.4E-6 | 378 | 398 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010080 | Biological Process | regulation of floral meristem growth | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0035265 | Biological Process | organ growth | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 523 aa Download sequence Send to blast |
MAPASNWLSF SLMSPMEMLR SSESESPFVP YDTSSTASPH YFLDNFYANG WTNPKSQGFF 60 ADGEDHNNQS KQGHADSTNL TSFIDPQSHH QPIPKLEDFL GDSSSMVRYS DSQTETQDSS 120 SLTHMYDQSS AYFGDHQQQQ DLKAITGFQA FSTNSGSEVE DSATMARTTQ LAGGEFTGHS 180 IESNGNELGG GFSSCGTNAL SLGVTTQSSS QSKPIVPADS DGSKKIADTF GQRTSIYRGV 240 TRHRWTGRYE AHLWDNSCRR EGQARKGRQV YLGGYDKEDK AARAYDLAAL KYWGPTATTN 300 FPVSTYSKEL EDMKHVTKQE FIASLRRKSS GFSRGASIYR GVTRHHQQGR WQARIGRVAG 360 NKDLYLGTFA TEEEAAEAYD IAAIKFRGIN AVTNFEMNRY DVEAIGKSSL PVGGAAKRLK 420 LSLEAEEQKP SVNHDQQRPQ CSSGSNNINF TSMQPAVQSI PCGIPYDAAA ATAYYHHNLF 480 QQFQPNYYGA CESSGLTPSI ATQMSMMPQQ PAEFYIWPHH QSN |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00504 | DAP | Transfer from AT5G10510 | Download |
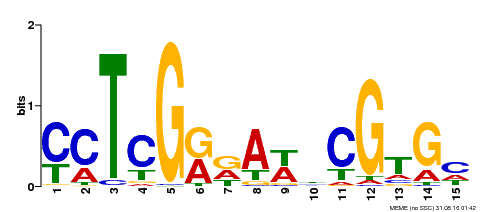
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_008236260.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008236260.1 | 0.0 | PREDICTED: AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Swissprot | Q52QU2 | 0.0 | AIL6_ARATH; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| TrEMBL | A0A251NT74 | 0.0 | A0A251NT74_PRUPE; Uncharacterized protein | ||||
| STRING | XP_008236260.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4810 | 31 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10510.2 | 0.0 | AINTEGUMENTA-like 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



