 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_004515250.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Cicereae; Cicer
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 459aa MW: 51024.8 Da PI: 6.3018 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.9 | 3.6e-16 | 61 | 105 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT E++++vda k++G g W+ I +++g ++t+ q++s+ qk+
XP_004515250.1 61 REKWTDKEHQKFVDALKLYGRG-WRQIEEHIG-TKTAVQIRSHAQKF 105
789*******************.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.84E-15 | 55 | 109 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.487 | 56 | 110 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.1E-8 | 58 | 109 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.8E-15 | 59 | 108 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 2.2E-11 | 60 | 108 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.2E-13 | 61 | 104 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.54E-9 | 63 | 106 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 459 aa Download sequence Send to blast |
MEIKNQVEGT KSTIIETESK CHSEGGEQPE NVAKSQDIPS VGNGNNLTPK VRKPYTITKQ 60 REKWTDKEHQ KFVDALKLYG RGWRQIEEHI GTKTAVQIRS HAQKFFSKVV REHDGSAESS 120 IQPIVIPPPR PKRKPLHPYP RKSVDSIKGQ PVPNKSETSP SVNLSVAEND TQSPTSVLSA 180 FASEAFGSAT FSEQTNRCLS PNSCTTETHP INLSPVEKEN DCMTSKPSEE GEKESLATVP 240 LSTDSKPLIC MKSEISSSQE TQCFKEDAAN KQHITSIKLF GRTVSMIDNQ KPMKEDDDDN 300 TKPITIKSDD ETNNVENEKV GQEGISGQLE TQLSLTMCNG MENPNENQCV GECAADVSRW 360 SLYQGLPAVN FKTSCNHQIL NPVPLRPFLK VRTREEESSC TGSNTESVCD NTVDSQTQKH 420 HQKSGRGFVP YKRCLAERDE NSLIVAFEER EGQRARVCS |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
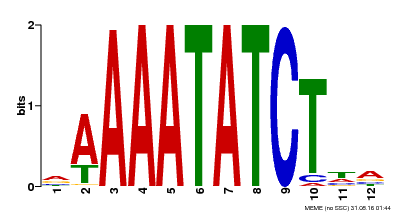
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_004515250.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004515250.1 | 0.0 | protein REVEILLE 7-like | ||||
| TrEMBL | A0A1S2Z547 | 0.0 | A0A1S2Z547_CICAR; protein REVEILLE 7-like | ||||
| STRING | XP_004515250.1 | 0.0 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7313 | 32 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.1 | 5e-57 | MYB_related family protein | ||||



