 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi0262s00020.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 224aa MW: 25598.9 Da PI: 5.7704 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.5 | 2.6e-17 | 5 | 50 | 3 | 48 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+WT+eEd ll+++++q+G g W++++ g++R++k+c++rw++yl
Vradi0262s00020.1 5 AWTEEEDHLLKKCIQQYGEGKWHRVPLLAGLNRCRKSCRLRWLNYL 50
7*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 49.2 | 1.2e-15 | 56 | 100 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rg++ +eE e +++++k+lG++ W++Ia +++ gRt++++k++w+ +
Vradi0262s00020.1 56 RGNFAEEEVEMIIKLHKLLGNR-WSLIAGRLP-GRTANDVKNYWNCH 100
8999******************.*********.***********977 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.777 | 1 | 50 | IPR017930 | Myb domain |
| SMART | SM00717 | 5.8E-13 | 2 | 52 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-21 | 5 | 57 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 7.7E-28 | 5 | 97 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 1.1E-15 | 5 | 50 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.65E-10 | 5 | 50 | No hit | No description |
| PROSITE profile | PS51294 | 24.633 | 51 | 105 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.8E-15 | 55 | 103 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.9E-13 | 56 | 98 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.07E-10 | 58 | 101 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.2E-24 | 58 | 105 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0031540 | Biological Process | regulation of anthocyanin biosynthetic process | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 224 aa Download sequence Send to blast |
MGGVAWTEEE DHLLKKCIQQ YGEGKWHRVP LLAGLNRCRK SCRLRWLNYL RPNIKRGNFA 60 EEEVEMIIKL HKLLGNRWSL IAGRLPGRTA NDVKNYWNCH LSKRLNAQEA EDKGGSRDVE 120 VIRPQARNMG SSSVKKRSEG VPPREKAVEQ ESSMSSLTFD ADGQNQVVES QEENIYACLD 180 QQGIVNEYSM DFQLEGFEDM MKGEGSSSQW DWGDLLLDMD LYK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 5e-22 | 6 | 107 | 10 | 110 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator, when associated with BHLH002/EGL3/MYC146, BHLH012/MYC1, or BHLH042/TT8. {ECO:0000269|PubMed:15361138}. | |||||
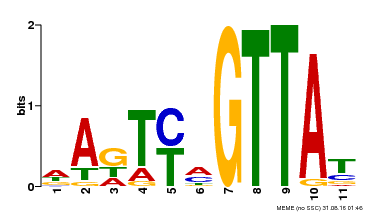
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00213 | DAP | Transfer from AT1G66370 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi0262s00020.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015037 | 0.0 | AP015037.1 Vigna angularis var. angularis DNA, chromosome 4, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014492680.1 | 1e-166 | transcription factor MYB113-like | ||||
| Swissprot | Q9FNV9 | 6e-55 | MY113_ARATH; Transcription factor MYB113 | ||||
| TrEMBL | A0A1S3TFY7 | 1e-164 | A0A1S3TFY7_VIGRR; transcription factor MYB113-like | ||||
| STRING | XP_007140451.1 | 1e-151 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF31 | 34 | 817 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G66370.1 | 1e-57 | myb domain protein 113 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



