 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Vradi01g01730.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Vigna
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 320aa MW: 35110.4 Da PI: 8.3466 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.5 | 1.3e-17 | 13 | 60 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +lv +++ +G+g+Wk+++ g+ R+ k+c++rw +yl
Vradi01g01730.1 13 KGPWTPEEDIILVSYIQDHGPGNWKAVPTNTGLSRCSKSCRLRWTNYL 60
79********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 51.6 | 2.1e-16 | 66 | 111 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T++E++ ++++ ++lG++ W++Ia++++ Rt++++k++w++yl
Vradi01g01730.1 66 RGNFTEQEEKMIIHLQALLGNR-WAAIAAYLP-QRTDNDIKNYWNTYL 111
89********************.*********.*************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 25.452 | 8 | 64 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 9.9E-31 | 10 | 107 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.9E-15 | 12 | 62 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.3E-16 | 13 | 60 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.5E-24 | 14 | 67 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.03E-11 | 15 | 60 | No hit | No description |
| SMART | SM00717 | 7.3E-16 | 65 | 113 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 20.864 | 65 | 115 | IPR017930 | Myb domain |
| Pfam | PF00249 | 9.0E-16 | 66 | 111 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.39E-11 | 68 | 111 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 5.2E-25 | 68 | 116 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001666 | Biological Process | response to hypoxia | ||||
| GO:0009617 | Biological Process | response to bacterium | ||||
| GO:0009626 | Biological Process | plant-type hypersensitive response | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0042761 | Biological Process | very long-chain fatty acid biosynthetic process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 320 aa Download sequence Send to blast |
MGRPPCCDKG VKKGPWTPEE DIILVSYIQD HGPGNWKAVP TNTGLSRCSK SCRLRWTNYL 60 RPGIKRGNFT EQEEKMIIHL QALLGNRWAA IAAYLPQRTD NDIKNYWNTY LKKKLNKLEA 120 SGSEGSLGHV GVSVSQSMSR GQWERRLQTD IHMAKKALSE ALSPKKTSST FLAELNSNRS 180 NDNSTGSFYN STKPTPSISY ASSADNIARL LKGWMKSTPK RAATQNSIIN LACAGADTAS 240 SEGTTKGSST SVELSETFES LFGYESLESS NSEFSPSLSP EGTLFQDESK PDITGIDVPF 300 SLLEKWLLDD SGCQEKLSF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 1e-26 | 13 | 115 | 27 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds specifically to the DNA sequence 5'-AACAAAC-3' (PubMed:19170933). Acts as a positive regulator of hypersensitive cell death (PubMed:10571865, PubMed:12119395). Acts as a positive regulator of salicylic acid synthesis (PubMed:16730712). Regulates very-long-chain fatty acid biosynthesis (PubMed:18326828). Acts cooperatively with BZR2 to promote expression of a subset of brassinosteroids target genes (PubMed:19170933). Transcriptional activity and hypersensitive response control negatively regulated by PLA2-ALPHA and by the Xanthomonas type III effector XopD (AC G9L9K6) (PubMed:20696912, PubMed:21917550). Involved in the regulation of abscisic acid (ABA) signaling (PubMed:22814374). Increased levels of MYB30 can accelerate flowering both in long and short days through the regulation of FT (PubMed:24587042). {ECO:0000269|PubMed:10571865, ECO:0000269|PubMed:12119395, ECO:0000269|PubMed:16730712, ECO:0000269|PubMed:18326828, ECO:0000269|PubMed:19170933, ECO:0000269|PubMed:20696912, ECO:0000269|PubMed:21917550, ECO:0000269|PubMed:22814374, ECO:0000269|PubMed:24587042}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00386 | DAP | Transfer from AT3G28910 | Download |
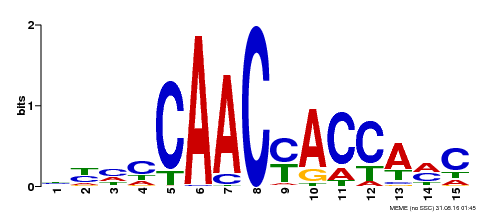
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Vradi01g01730.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated during hypersensitive response, but no expression detected during compatible interaction with pathogens (PubMed:10571865). Specifically induced in the inoculated zone 4 hours post pathogen infection (PubMed:20696912). Up-regulated by jasmonic acid and salicylic acid (PubMed:16463103, PubMed:16730712). Transcriptionally regulated by BZR2 (PubMed:19170933). {ECO:0000269|PubMed:10571865, ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:16730712, ECO:0000269|PubMed:19170933, ECO:0000269|PubMed:20696912}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015044 | 0.0 | AP015044.1 Vigna angularis var. angularis DNA, chromosome 11, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014506098.1 | 0.0 | transcription factor MYB30 | ||||
| Swissprot | Q9SCU7 | 1e-112 | MYB30_ARATH; Transcription factor MYB30 | ||||
| TrEMBL | A0A1S3UJ93 | 0.0 | A0A1S3UJ93_VIGRR; transcription factor MYB30 | ||||
| STRING | XP_007151227.1 | 0.0 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1016 | 34 | 115 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G28910.1 | 1e-109 | myb domain protein 30 | ||||



