 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Traes_6DS_2B2F9C290.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Pooideae; Triticodae; Triticeae; Triticinae; Triticum
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 870aa MW: 97606.6 Da PI: 7.2628 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.5 | 1.2e-05 | 557 | 579 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ Cg +F+ + +Lk+H+++H
Traes_6DS_2B2F9C290.2 557 HSCEECGACFRKPAHLKQHMQSH 579
79*******************99 PP
| |||||||
| 2 | zf-C2H2 | 20.2 | 1.6e-06 | 585 | 609 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC+ s+ r+++L+rH+ tH
Traes_6DS_2B2F9C290.2 585 FACPleDCPFSYIRRDHLNRHMLTH 609
78**********************9 PP
| |||||||
| 3 | zf-C2H2 | 15.7 | 4.2e-05 | 614 | 639 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++Cp Cgk+Fs k n +rH + H
Traes_6DS_2B2F9C290.2 614 FTCPleGCGKRFSIKANIQRHVKEmH 639
89********************9988 PP
| |||||||
| 4 | zf-C2H2 | 15.4 | 5.5e-05 | 658 | 680 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
C+ C+k F+ +s+Lk+H +H
Traes_6DS_2B2F9C290.2 658 CQeeGCKKAFKYPSQLKKHEESH 680
77779**************9988 PP
| |||||||
| 5 | zf-C2H2 | 13.3 | 0.00026 | 747 | 771 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C sFs+ksnL+ H++ H
Traes_6DS_2B2F9C290.2 747 KCSfdGCECSFSNKSNLNMHMKAcH 771
799999****************955 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51375 | 5.426 | 35 | 65 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.451 | 66 | 100 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 1.3E-5 | 66 | 98 | IPR011990 | Tetratricopeptide-like helical domain |
| Pfam | PF01535 | 0.77 | 68 | 92 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.627 | 136 | 170 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 2.5E-4 | 141 | 163 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 1.8E-4 | 141 | 164 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.0078 | 169 | 195 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 9.986 | 198 | 232 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 2.1E-4 | 200 | 233 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 4.1E-4 | 200 | 229 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 7.739 | 268 | 298 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.0075 | 273 | 294 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 1.3E-5 | 298 | 356 | IPR011990 | Tetratricopeptide-like helical domain |
| Pfam | PF13041 | 7.1E-11 | 298 | 346 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 12.233 | 299 | 333 | IPR002885 | Pentatricopeptide repeat |
| TIGRFAMs | TIGR00756 | 2.2E-8 | 301 | 335 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 8.572 | 334 | 364 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 6.138 | 370 | 400 | IPR002885 | Pentatricopeptide repeat |
| Pfam | PF01535 | 0.4 | 373 | 397 | IPR002885 | Pentatricopeptide repeat |
| Gene3D | G3DSA:1.25.40.10 | 1.3E-5 | 402 | 464 | IPR011990 | Tetratricopeptide-like helical domain |
| PROSITE profile | PS51375 | 5.327 | 402 | 432 | IPR002885 | Pentatricopeptide repeat |
| PROSITE profile | PS51375 | 7.322 | 436 | 470 | IPR002885 | Pentatricopeptide repeat |
| SMART | SM00355 | 42 | 511 | 534 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.277 | 557 | 584 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.008 | 557 | 579 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.6E-7 | 557 | 584 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 559 | 579 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 9.54E-11 | 570 | 611 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 9.9E-8 | 585 | 611 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 7.2E-4 | 585 | 609 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.536 | 585 | 614 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 587 | 609 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.19E-8 | 595 | 646 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 4.1E-6 | 612 | 639 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 12.133 | 614 | 644 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 4.0E-4 | 614 | 639 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 616 | 639 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.245 | 656 | 681 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.039 | 656 | 680 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 8.6E-6 | 658 | 680 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 658 | 680 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 5 | 688 | 713 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 690 | 713 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 18 | 716 | 737 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 1.59E-8 | 731 | 788 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.9E-5 | 731 | 768 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.99 | 746 | 776 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0027 | 746 | 771 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 748 | 771 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 8.642 | 777 | 803 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 2.2 | 777 | 803 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 779 | 803 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 870 aa Download sequence Send to blast |
MNRSVCHHLL AHCRSLRELE RIHARAVTHG LHPSNQSISC TLFRRYADFG RPAEARRLFG 60 EIPCPDLVSY TSLMSLHLQL SQHHEAMSAF SCAVASGHRP DGFVVVGALS ASGGAGDLRA 120 GLGVHGLIFR CGLGSELVVG NALVDMYCRC GRFEGVLGVF DEMAVKDEVT WGSMLHGYMK 180 CVGVDAALSF FDQMPVKSTV SWTALVTGHV QAKQPIRALE IFGRMVLEGH RPTHVTIVGV 240 LSACADIGSL DLGRVIHGYG SKFNISTNII VSNALMDMYA KSGSIEMAFT VFEEVPLKDA 300 FTWTTMISSF TVQGDGIKAL ELFEDMLRSG IVPNSVTFVS VLSACSHAGL IQQGRELFDN 360 MRRIYNIKPQ LEHYGCIVDL LGRGGLLEEA EALIYDMDVE PDTVIWRSLL SACLVHGNDR 420 LAEIAGQEII KREPGDDGVY VLLWNIYASS DRWKEALDIR KQMLTRKIFK KPGCSWIEVD 480 GGVHEFLMSS GDGIDGDGKS EETHSKDIRR IKCEFCTVVR SKMYLIRAHM VAQHQDELDS 540 SEIYNSNGEK VVHGVGHSCE ECGACFRKPA HLKQHMQSHS KERSFACPLE DCPFSYIRRD 600 HLNRHMLTHE GKLFTCPLEG CGKRFSIKAN IQRHVKEMHE EEHECKMDDT KSNQQVICQE 660 EGCKKAFKYP SQLKKHEESH AKLDYVEVVC CEPGCLKTFT NAKCLRAHNQ SCHLYTQCDI 720 CGEKHLRKNI KRHLRAHHKV HSTERLKCSF DGCECSFSNK SNLNMHMKAC HDQVRPFECR 780 VAGCGKTFPY KHVRDNHEKS SCHVYVEGDY EEMDEQLRSR PRGGRKRAAV TVETLTRKRV 840 TIPSEASSLE DGAEYMRWML SDGDGSNHAE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5iww_D | 3e-56 | 9 | 412 | 4 | 330 | PLS9-PPR |
| Search in ModeBase | ||||||
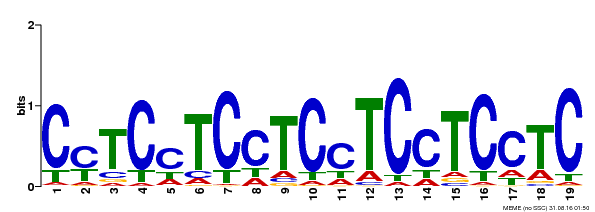
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK374527 | 0.0 | AK374527.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv3068C12. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020181266.1 | 0.0 | putative pentatricopeptide repeat-containing protein At5g59200, chloroplastic | ||||
| TrEMBL | A0A453MVM8 | 0.0 | A0A453MVM8_AEGTS; Uncharacterized protein | ||||
| STRING | Traes_6DS_2B2F9C290.2 | 0.0 | (Triticum aestivum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 5e-90 | transcription factor IIIA | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



