 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp7g13320 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 812aa MW: 93780.9 Da PI: 6.5688 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 91.6 | 1.1e-28 | 68 | 157 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+fY+eYAk++GF++++++s++sk+++ +++++f Cs++g + e+++ ++++r++++++t+Cka+++vk++ dgkw ++++++eHnHel p
Tp7g13320 68 SFYQEYAKSMGFTTSIKNSRRSKKTKDFIDAKFACSRYGVTPESESG-SSSSRRSSVKKTDCKASMHVKRRPDGKWIIHEFVKEHNHELLP 157
5****************************************999998.89999999********************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 3.2E-26 | 68 | 157 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.4E-24 | 278 | 371 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 8.803 | 558 | 594 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 4.2E-5 | 560 | 592 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 3.9E-5 | 569 | 596 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 812 aa Download sequence Send to blast |
MDLQENNNQV SDAGDDNMVD VVVESHNNNR DVGIVVDDLR TGDLGFSGDL DLEPRNGIDF 60 DTHEAAYSFY QEYAKSMGFT TSIKNSRRSK KTKDFIDAKF ACSRYGVTPE SESGSSSSRR 120 SSVKKTDCKA SMHVKRRPDG KWIIHEFVKE HNHELLPALA YHFRIQRNVK LAEKNNIDIL 180 HAVSERTRKM YVEMSRQSGG YKNIRLLPHT DVNSQVDKDR YLALEEGDSQ VLLEYFKRIK 240 KENPKFFYAI DLNKEQRLRN LFWADAKSRD DYMSFNDVVS FDTTYVKFND KLPLGLFIGV 300 NHHSQPMLLG CALVADESKE TFVWLIKTWL RAMSGGAPKV ILTDQDKLLM TAVSELLPNT 360 RHCFALWHVL EKIPEYFSHV MKRHENFLQK FNKCIFRSWT DDQFDMRWWK MVSQFGLEDD 420 EWLLWLHEHR QKWVPTFMSD VFLAGMSTSQ RSESVNSFFD KYVHKKITLK EFLRQYGAIL 480 QNRYEEESVA DFDTCHKQPA LKSPSPWEKQ MATTYTHTIF KKFQVEVLGV VACHPRKEKE 540 DENIVTFRVQ DCEKDDEFLV TWSKTKSELC CFCHMFEYKG FLCRHALMIL QICGFASIPP 600 RYILKRWTKD AKSGVLAGEG ADQTQARVQR YNDLCSRAVE LSEEGCVSEE NYNIVLRTLV 660 ETLKNCVDLN NARNNITESN SQLNSGAHEE ENQGQPEASQ MLESQQSLQP MENISSEGMS 720 ISGYYGPQQN VQGLLNLMEP PHEGYYVNQR TIQGLGQLNS IAPVHDSFFT NQQAMPGMVG 780 QIDFRPPPNF TYNLQDEHLR SAQLPGSSSR HL |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
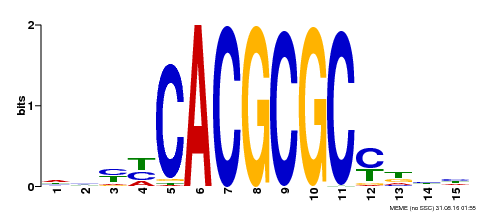
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp7g13320 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF159587 | 0.0 | AF159587.1 Arabidopsis thaliana far-red impaired response protein (FAR1) mRNA, complete cds. | |||
| GenBank | CP002687 | 0.0 | CP002687.1 Arabidopsis thaliana chromosome 4 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006414566.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | A0A1J3JL92 | 0.0 | A0A1J3JL92_NOCCA; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| STRING | XP_006414566.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||



