 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp6g14420 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Brassicaceae incertae sedis; Schrenkiella
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 699aa MW: 78110.9 Da PI: 6.7741 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.2 | 6.9e-19 | 53 | 107 | 2 | 56 |
T--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 2 rkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k +++t++q++eLe++F+ +++p++++r eL +kl L+ rq+k+WFqNrR+++k
Tp6g14420 53 TKYRRHTSYQIQELESFFKACPHPNEKQRLELGSKLALESRQIKFWFQNRRTQMK 107
678899**********************************************999 PP
| |||||||
| 2 | START | 170.7 | 9.5e-54 | 225 | 442 | 5 | 206 |
HHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS...SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECTT...... CS
START 5 eaaqelvkkalaeepgWvkss....esengdevlqkfeeskv..dsgealrasgvvdmvlallveellddkeqWdetla....kaetlevissg...... 88
a++el+k+a+++ p+W k s es n de++++f+ k + e ++++g+v ++ +lve+l+d++ +W e+++ a+t+evis+g
Tp6g14420 225 VAMDELIKLAELDNPLWIKCSksekESMNHDEYRSIFTGRKHpgFIAEGSKETGFVLINSLTLVEILMDTN-RWAEMCEcivaVASTIEVISNGsggstn 323
589******************99999999********777768899999**********************.**************************** PP
EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHH CS
START 89 galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvks 187
g lqlm ae+q++splvp R + f+Ry++q+++g w++vdvS d +++ ++ +s+ ++lpSg++i++ +ng skvtw+eh +++ +++h l+++l++s
Tp6g14420 324 GSLQLMEAEFQVISPLVPiRQVKFLRYCKQHEDGLWAVVDVSYDMNKEDENLKSYDGFRRLPSGCIIQDIGNGCSKVTWIEHSEYERNQIHPLYKQLLSS 423
*************************************************9988888788***************************************** PP
HHHHHHHHHHHHTXXXXXX CS
START 188 glaegaktwvatlqrqcek 206
++ ga +w+atlqrqce+
Tp6g14420 424 SVGLGATKWLATLQRQCES 442
*****************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.88E-18 | 36 | 109 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 6.8E-20 | 38 | 109 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.989 | 49 | 109 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.2E-15 | 51 | 113 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.3E-16 | 53 | 107 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.16E-14 | 56 | 109 | No hit | No description |
| PROSITE profile | PS50848 | 46.093 | 212 | 445 | IPR002913 | START domain |
| CDD | cd08875 | 4.39E-108 | 217 | 441 | No hit | No description |
| SuperFamily | SSF55961 | 7.83E-30 | 217 | 443 | No hit | No description |
| SMART | SM00234 | 1.4E-38 | 221 | 442 | IPR002913 | START domain |
| Pfam | PF01852 | 4.0E-47 | 225 | 442 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.18E-15 | 473 | 692 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 699 aa Download sequence Send to blast |
MNGDLDLDMS RGDLNPSFLR RFRDDEFESD SFDAMSGGDD KDEQRPKKKK KKTKYRRHTS 60 YQIQELESFF KACPHPNEKQ RLELGSKLAL ESRQIKFWFQ NRRTQMKTQL ERHENAILRQ 120 ENEKLRVENG VLKEAMRNPI CNNCGGAVIP DEVSYEQQQL RIENAKLKYE LDKLCALANR 180 FIGGSISFEQ PSTGGNGSQD FSTRHGFTRG TLMCESSMFL DLAMVAMDEL IKLAELDNPL 240 WIKCSKSEKE SMNHDEYRSI FTGRKHPGFI AEGSKETGFV LINSLTLVEI LMDTNRWAEM 300 CECIVAVAST IEVISNGSGG STNGSLQLME AEFQVISPLV PIRQVKFLRY CKQHEDGLWA 360 VVDVSYDMNK EDENLKSYDG FRRLPSGCII QDIGNGCSKV TWIEHSEYER NQIHPLYKQL 420 LSSSVGLGAT KWLATLQRQC ESYTTLLSSQ DHTGLSLAGT KSTLTLAQRM KCNFYSGITA 480 SPIHKWDKLV AENLGQDTRI LTRKSLQPSG IVLSAATSMW LPVTQQRLFE FLCDSKCRNQ 540 WDILANGASM ENMLLIPKGQ HEGRCVSLLS PAGKHQNESS MLILQETWND ASGALVVYAP 600 VDVPSMNIVM SGGDSANMAL LPSGFSILPD GSSSSWSDQT STNGSLVNHE SKGCLLTLGF 660 QIIVNSKPTA KLNMESVQTV NNLIACTIHK IKAALGIPA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds to the DNA sequence 5'-GCATTAAATGC-3'. {ECO:0000269|PubMed:16778018}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00559 | DAP | Transfer from AT5G52170 | Download |
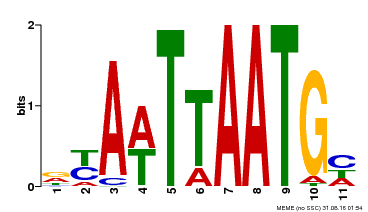
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp6g14420 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009106897.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein HDG7 | ||||
| Refseq | XP_013686466.1 | 0.0 | homeobox-leucine zipper protein HDG7-like | ||||
| Swissprot | Q9LTK3 | 0.0 | HDG7_ARATH; Homeobox-leucine zipper protein HDG7 | ||||
| TrEMBL | A0A3P6CY22 | 0.0 | A0A3P6CY22_BRACM; Uncharacterized protein | ||||
| STRING | Bra028295.1-P | 0.0 | (Brassica rapa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52170.1 | 0.0 | homeodomain GLABROUS 7 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



