 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Tp57577_TGAC_v2_mRNA10715 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Trifolieae; Trifolium
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 296aa MW: 34094.8 Da PI: 6.9369 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 58.6 | 1.1e-18 | 98 | 150 | 4 | 56 |
-SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 4 RttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+ +++ eq+++Le+ Fe +++ +++ +LAk lgL+ rqV +WFqNrRa++k
Tp57577_TGAC_v2_mRNA10715 98 KKRLSLEQVKALEKSFEIGNKLEPDRKMQLAKALGLQPRQVAIWFQNRRARWK 150
556899**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 129.7 | 1.2e-41 | 96 | 188 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekev 84
ekk+rls eqvk+LE+sFe +kLep+rK++la++Lglqprqva+WFqnrRAR+ktkqlEk+ye+Lk+++++lk++n++L++++
Tp57577_TGAC_v2_mRNA10715 96 EKKKRLSLEQVKALEKSFEIGNKLEPDRKMQLAKALGLQPRQVAIWFQNRRARWKTKQLEKEYEVLKKQFESLKADNDSLKAQN 179
69********************************************************************************** PP
HD-ZIP_I/II 85 eeLreelke 93
++L++el++
Tp57577_TGAC_v2_mRNA10715 180 HKLHAELQT 188
*****9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.11E-19 | 89 | 154 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.022 | 92 | 152 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.6E-19 | 95 | 156 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 9.40E-16 | 97 | 153 | No hit | No description |
| Pfam | PF00046 | 7.8E-16 | 98 | 150 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 7.2E-20 | 104 | 151 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 8.4E-6 | 123 | 132 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 127 | 150 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 8.4E-6 | 132 | 148 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 2.6E-15 | 152 | 192 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 296 aa Download sequence Send to blast |
MAFPPSHTFI FQTHEDHHHD HLHSSSSSLN SFPSFPPHQH FQGSGNHGGA SYMMKRSMSF 60 SGIENNHNHM NNKCDELMHG DEDQLSDEEG YSQLGEKKKR LSLEQVKALE KSFEIGNKLE 120 PDRKMQLAKA LGLQPRQVAI WFQNRRARWK TKQLEKEYEV LKKQFESLKA DNDSLKAQNH 180 KLHAELQTLK NRDCFENGTI SLKKENECSW SNGSDNSSDM NLDLSRTQIM SSPLSSQNGK 240 TLLQTSLKPT SMTQLLQSST RSDLQDESFC NMFHNIDEQQ SLWPWTDHQQ QHHFQ* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 144 | 152 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00322 | DAP | Transfer from AT3G01220 | Download |
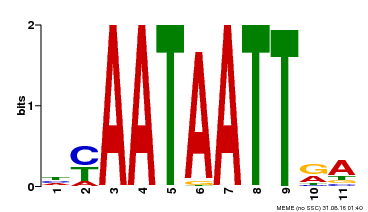
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Tp57577_TGAC_v2_mRNA10715 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:12644682}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT148482 | 0.0 | BT148482.1 Medicago truncatula clone JCVI-FLMt-17G6 unknown mRNA. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_013450953.1 | 1e-178 | homeobox-leucine zipper protein HAT7 | ||||
| Swissprot | Q8LAT0 | 5e-81 | ATB20_ARATH; Homeobox-leucine zipper protein ATHB-20 | ||||
| TrEMBL | A0A2K3PC66 | 0.0 | A0A2K3PC66_TRIPR; Homeobox-leucine zipper protein HAT7-like | ||||
| STRING | XP_004489381.1 | 1e-158 | (Cicer arietinum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1271 | 33 | 106 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G15150.1 | 3e-69 | homeobox 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Tp57577_TGAC_v2_mRNA10715 |



