 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10024410m | ||||||||
| Common Name | EUTSA_v10024410mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 829aa MW: 95425.3 Da PI: 7.7137 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 91.2 | 1.4e-28 | 66 | 156 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHelap 91
+fY+eYAk++GF++++++s++sk+++ +++++f Cs++g + e+++ +++++r++++++t+Cka+++vk++ dgkw ++++ ++HnHel p
Thhalv10024410m 66 SFYQEYAKSMGFTTSIKNSRRSKKTKDFIDAKFACSRYGVTPESESGSSSSSRRSSVKKTDCKASMHVKRRPDGKWIIHEFIKDHNHELLP 156
5******************************************999999***************************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 3.1E-26 | 66 | 156 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 5.7E-26 | 277 | 370 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9 | 557 | 593 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 2.6E-5 | 557 | 591 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 3.9E-5 | 568 | 595 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 829 aa Download sequence Send to blast |
MDLQVSNQVS DAGHENMVGV VVESHSNRDV GIVVDDLRTG GLGFSGDLDL EPRNGIDFDT 60 HEAAYSFYQE YAKSMGFTTS IKNSRRSKKT KDFIDAKFAC SRYGVTPESE SGSSSSSRRS 120 SVKKTDCKAS MHVKRRPDGK WIIHEFIKDH NHELLPALAY HFRIQRNVKL AEKNNIDILH 180 AVSERTRKMY VEMSRQSGGY KNIGLVPQTD VNSQVDKGRY LALEEGDSQL LLEYFKRIKK 240 ENPKFFYAID LNEEQRLRNL FWADAKSRDD YLSFNDVVSF DTTYVKFNDK LPLALFIGVN 300 HHSQPMLLGC ALVADESKET FVWLIKTWLR AMGGRAPKVI LTDQDKLLMA AVSELLPNTR 360 HCFALWHVLE KIPEYFSHVM KRHENFLRKF NKCIFRSWTG DQFEMRWWKM VSRFGLEEDE 420 WLQWLHEHRQ KWVPTFMSDV FLAGMSTSQR SESVNSFFDK YIHKKITLKE FLRQYGAILQ 480 NRYEEESVAD FDTCHKQPAL KSPSPWEKQM ATTYTHTIFK KFQVEVLGVV ACHPRKEKED 540 ENMATFRVQD CEKDDEYFVT WSKTKSELCC FCHMFEYKGF LCRHALMILQ ICGFASIPPR 600 YILKRWTKDA KSGVLAGEGA DQIQTRAQRY NDLCRRAIEL SEEGCVSEEN YNIVLRTLVE 660 TLKNCVDLNN SRSNITESNS QLNNGAHEEE NQVMASLKAT KKKTVVRKRK GQPEASQMLE 720 SQQSLQPMEI ISSEGLSING FYGPQQNVQG LLNLMEPPHE GYYVNQRTIQ GLGQLNSIAP 780 VQDSFFTNQQ GMPGLGQIDF RPPPNFTYSL QDEHLRSAQL PGSSSREL* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
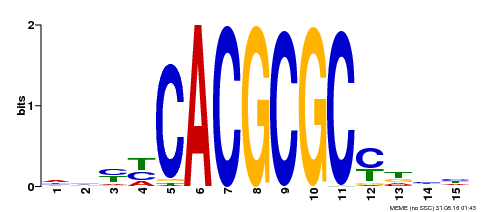
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10024410m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. {ECO:0000269|PubMed:18033885}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF159587 | 0.0 | AF159587.1 Arabidopsis thaliana far-red impaired response protein (FAR1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006414566.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| Swissprot | Q9SWG3 | 0.0 | FAR1_ARATH; Protein FAR-RED IMPAIRED RESPONSE 1 | ||||
| TrEMBL | V4MHH4 | 0.0 | V4MHH4_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006414566.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM118 | 26 | 253 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 0.0 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10024410m |
| Entrez Gene | 18028864 |



