|
Plant Transcription
Factor Database
|
Transcription Factor Information
|
Basic
Information? help
Back to Top |
| TF ID |
Thhalv10019007m |
| Common Name | EUTSA_v10019007mg |
| Organism |
|
| Taxonomic ID |
|
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
| Family |
MYB_related |
| Protein Properties |
Length: 267aa MW: 29114.3 Da PI: 10.5053 |
| Description |
MYB_related family protein |
| Gene Model |
| Gene Model ID |
Type |
Source |
Coding Sequence |
| Thhalv10019007m | genome | JGI | View CDS |
|
| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | Myb_DNA-binding | 44.1 | 4.7e-14 | 103 | 147 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT++E+ l++ + ++ G+g+W+ I+r + k+Rt+ q+ s+ qky
Thhalv10019007m 103 PWTEDEHRLFLTGLHKVGKGDWRGISRNFVKTRTPTQVASHAQKY 147
8*******************************************9 PP
|
| Gene Ontology ? help Back to Top |
| GO Term |
GO Category |
GO Description |
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated |
| GO:0009651 | Biological Process | response to salt stress |
| GO:0009723 | Biological Process | response to ethylene |
| GO:0009733 | Biological Process | response to auxin |
| GO:0009737 | Biological Process | response to abscisic acid |
| GO:0009739 | Biological Process | response to gibberellin |
| GO:0009751 | Biological Process | response to salicylic acid |
| GO:0009753 | Biological Process | response to jasmonic acid |
| GO:0031540 | Biological Process | regulation of anthocyanin biosynthetic process |
| GO:0046686 | Biological Process | response to cadmium ion |
| GO:0080167 | Biological Process | response to karrikin |
| GO:0005634 | Cellular Component | nucleus |
| GO:0003677 | Molecular Function | DNA binding |
| GO:0003682 | Molecular Function | chromatin binding |
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding |
| GO:0008270 | Molecular Function | zinc ion binding |
| Sequence ? help Back to Top |
| Protein Sequence Length: 267 aa
Download sequence Send
to blast |
MSRSCSQCGN NGHNSRTCPT EISAAAGEDG GGGGGENKGI MLFGVRVTEA SSSSFRKSVS 60
MNNLSQFDHA AHDSNPVDDG GYASDDVVHA SSVRNRERKR GTPWTEDEHR LFLTGLHKVG 120
KGDWRGISRN FVKTRTPTQV ASHAQKYFLR RTNQNRRRRR SSLFDITPDS FIGTSTEEKN 180
QSQTPLERMK HIRPVPIPIP IPPSRKMADL NLNQKTSAPA AAEMFPLSLN LQVKTTSSSS 240
SSNEQKTRGS AFDTMSNNGD SIMGMA*
|
| Functional Description ? help
Back to Top |
| Source |
Description |
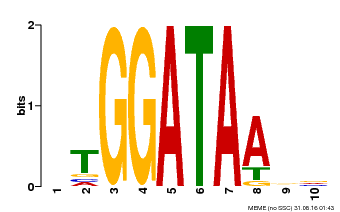
| UniProt | Transcription repressor that binds to 5'-TATCCA-3' elements in gene promoters. Contributes to the sugar-repressed transcription of promoters containing SRS or 5'-TATCCA-3' elements. Transcription repressor involved in a cold stress response pathway that confers cold tolerance. Suppresses the DREB1-dependent signaling pathway under prolonged cold stress. DREB1 responds quickly and transiently while MYBS3 responds slowly to cold stress. They may act sequentially and complementarily for adaptation to short- and long-term cold stress (PubMed:20130099). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:20130099}. |
| Regulation -- Description ? help
Back to Top |
| Source |
Description |
| UniProt | INDUCTION: Repressed by sucrose and gibberellic acid (GA) (PubMed:12172034). Induced by cold stress in roots and shoots. Induced by salt stress in shoots. Down-regulated by abscisic aci (ABA) in shoots (PubMed:20130099). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:20130099}. |
| Publications
? help Back to Top |
- Rice Chromosome 10 Sequencing Consortium
In-depth view of structure, activity, and evolution of rice chromosome 10.
Science, 2003. 300(5625): p. 1566-9
[PMID:12791992] - Kikuchi S, et al.
Collection, mapping, and annotation of over 28,000 cDNA clones from japonica rice.
Science, 2003. 301(5631): p. 376-9
[PMID:12869764] - Su CF, et al.
A novel MYBS3-dependent pathway confers cold tolerance in rice.
Plant Physiol., 2010. 153(1): p. 145-58
[PMID:20130099]
|