 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10004645m | ||||||||
| Common Name | EUTSA_v10004645mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 311aa MW: 34744.6 Da PI: 4.8293 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 60.8 | 2.1e-19 | 70 | 123 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++++eq+++Le+ Fe ++++ e++ +LA++lgL+ rqV +WFqNrRa++k
Thhalv10004645m 70 KKRRLSAEQVKALEKNFEIDNKLEPERKVKLAQELGLQPRQVAIWFQNRRARWK 123
56689************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 130.5 | 6.6e-42 | 69 | 161 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekkrrls+eqvk+LE++Fe ++kLeperKv+la+eLglqprqva+WFqnrRAR+ktkqlE+dy +Lk+++dalk+++++L++++++L +e+ke
Thhalv10004645m 69 EKKRRLSAEQVKALEKNFEIDNKLEPERKVKLAQELGLQPRQVAIWFQNRRARWKTKQLERDYGVLKSNFDALKRSRDSLQRDNDSLFAEIKE 161
69**************************************************************************************99987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 5.43E-20 | 60 | 127 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.977 | 65 | 125 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 9.1E-19 | 68 | 129 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 4.14E-18 | 70 | 126 | No hit | No description |
| Pfam | PF00046 | 1.3E-16 | 70 | 123 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 6.2E-21 | 71 | 132 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 7.7E-6 | 96 | 105 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 100 | 123 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 7.7E-6 | 105 | 121 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 6.4E-17 | 125 | 166 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 311 aa Download sequence Send to blast |
MKRSRGSSDS LSGFLPICHS ATDKQISPRP TTTGFLYSGA GDYSPMFDGL EDDGSLEDIG 60 VRHASAAAEK KRRLSAEQVK ALEKNFEIDN KLEPERKVKL AQELGLQPRQ VAIWFQNRRA 120 RWKTKQLERD YGVLKSNFDA LKRSRDSLQR DNDSLFAEIK ELRAKVNVEG PGGSNGNTSI 180 EEDGIAKRVE AMVNQSNKEV LQLSHRPPPP NIPTEVPTSE LTYEMFNIFP RTESFREDPA 240 DSSDSSAVLN EEYSPTRAEA AAATAVEMST MGCFSQFVKM EEHEDLFSGE EACKLFADNE 300 QWYCSGQWNS * |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 117 | 125 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
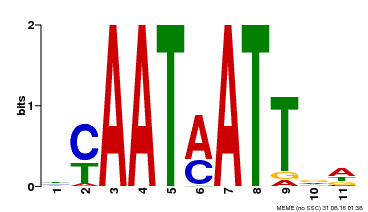
| UniProt | Probable transcription factor that acts as a positive regulator of ABA-responsiveness, mediating the inhibitory effect of ABA on growth during seedling establishment. Binds to the DNA sequence 5'-CAATNATTG-3'. {ECO:0000269|PubMed:12678559}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00587 | DAP | Transfer from AT5G65310 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10004645m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by abscisic acid (ABA) and by salt stress. {ECO:0000269|PubMed:12678559, ECO:0000269|PubMed:16055682}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353247 | 0.0 | AK353247.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-23-D07. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006394048.1 | 0.0 | homeobox-leucine zipper protein ATHB-5 | ||||
| Swissprot | P46667 | 1e-171 | ATHB5_ARATH; Homeobox-leucine zipper protein ATHB-5 | ||||
| TrEMBL | V4KJE3 | 0.0 | V4KJE3_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006394048.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM12986 | 15 | 26 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G65310.1 | 1e-163 | homeobox protein 5 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10004645m |
| Entrez Gene | 18013027 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



