 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10003641m | ||||||||
| Common Name | EUTSA_v10003641mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 855aa MW: 95229.6 Da PI: 6.8735 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 71.5 | 1.1e-22 | 164 | 265 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrs 88
f+k+lt sd++++g +++ +++a+e+ + + + +++l+ +d++ ++W++++i+r++++r++l++GW+ Fv++++L +gD ++F + +
Thhalv10003641m 164 FCKTLTASDTSTHGGFSVLRRHADEClppldmSRQ-PPTQELVAKDLHANEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RGE 254
99*****************************7333.34459************************************************..449 PP
EE..EEEEE-S CS
B3 89 efelvvkvfrk 99
++el+v+v+r+
Thhalv10003641m 255 NGELRVGVRRA 265
999*****996 PP
| |||||||
| 2 | Auxin_resp | 120.8 | 1e-39 | 290 | 372 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+st+++F+v+Y+Pr+s+seF+v+++++++++k+++s+GmRfkm+fe+e+++e+r++Gt+vg++d+dp+rW +SkWrsLk
Thhalv10003641m 290 AWHAISTGTMFTVYYKPRTSPSEFIVPFDQYMESVKNNYSIGMRFKMRFEGEEAPEQRFTGTIVGIEDSDPTRWAKSKWRSLK 372
89*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 3.01E-40 | 160 | 293 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.2E-40 | 160 | 279 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.45E-20 | 162 | 264 | No hit | No description |
| PROSITE profile | PS50863 | 11.42 | 164 | 266 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 4.1E-22 | 164 | 266 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.5E-20 | 164 | 265 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 7.7E-37 | 290 | 372 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 1.3E-10 | 715 | 813 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 1.08E-11 | 728 | 805 | No hit | No description |
| PROSITE profile | PS51745 | 25.174 | 728 | 812 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 855 aa Download sequence Send to blast |
MASSEVSVKG SRGGENFSSA GYSDPKETRN VSVVGEGQKS NSNRSVVAER AVDPEAALYR 60 ELWHACAGPL VTVPRQDDRV FYFPQGHIEQ VEASTNQAAE QQMPLYDLPS KLLCRVINVD 120 LKAEADTDEV YAQITLLPEP NQDENAIEKE TPPPPPPRFQ VHSFCKTLTA SDTSTHGGFS 180 VLRRHADECL PPLDMSRQPP TQELVAKDLH ANEWRFRHIF RGQPRRHLLQ SGWSVFVSSK 240 RLVAGDAFIF LRGENGELRV GVRRAMRQQG NVPSSVISSH SMHLGVLATA WHAISTGTMF 300 TVYYKPRTSP SEFIVPFDQY MESVKNNYSI GMRFKMRFEG EEAPEQRFTG TIVGIEDSDP 360 TRWAKSKWRS LKVRWDETSS IPRPDRVSPW KIEPALAPPA LSPVPMPRPK RPRSNIAPSS 420 PDSSMLQREG STKANMDPLP ASGLSRVLQG QEYSTLRTKH TESVECDVPE NSVVWQSSAD 480 DDKVDVVSAS RRCENWMSSG RHEPAPFTDL LSGFGANINP SFGQRIPFYD HSSSPSVPAK 540 KILSDHDGKF DFFANQWQMM HSGLSLKLHE SPKVSAPSDA SFQGRGNVKY GEYPVLHDLT 600 TENTGGNWPI RPRALNYFED VVHSQAREHV AKRPAVVQEE TTKARDGNCR LFGIPLVNNM 660 NGADSTMAQR NNLKDAAGLT QTAPPKVQDL SDQSKGSKST NDQREQGRPF QTNHPHPKDA 720 HTKTNSSRSC TKVHKQGIAL GRSVDLSKFQ NYEELIAELD RLFEFNGELM APKKDWLIVY 780 TDDENDMMLV GDDPWQEFCC MVRKIFIYTK EEVRKMNPGT LSCRSEEEAV VGEGSDAKDA 840 KSASNPSLSS AGNS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-180 | 57 | 394 | 18 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-180 | 57 | 394 | 18 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-180 | 57 | 394 | 18 | 354 | Auxin response factor 1 |
| 4ldx_A | 1e-180 | 57 | 394 | 18 | 354 | Auxin response factor 1 |
| 4ldx_B | 1e-180 | 57 | 394 | 18 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
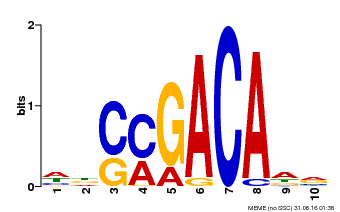
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10003641m |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353097 | 0.0 | AK353097.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-13-O23. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006394444.1 | 0.0 | auxin response factor 2 | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | V4K1E7 | 0.0 | V4K1E7_EUTSA; Auxin response factor | ||||
| STRING | XP_006394444.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5348 | 26 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10003641m |
| Entrez Gene | 18012800 |



