 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10001374m | ||||||||
| Common Name | EUTSA_v10001374mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 577aa MW: 64103.9 Da PI: 6.3118 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 43.7 | 5.1e-14 | 403 | 449 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr+++P+ + K++Ka+ L A+ YIk+Lq
Thhalv10001374m 403 NHVEAERQRREKLNQRFYALRSVVPNiS------KMDKASLLGDAISYIKELQ 449
799***********************66......******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50231 | 8.734 | 24 | 119 | IPR000772 | Ricin B, lectin domain |
| Pfam | PF14215 | 2.7E-51 | 48 | 239 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| SuperFamily | SSF47459 | 3.01E-19 | 398 | 463 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 17.936 | 399 | 448 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.60E-15 | 402 | 453 | No hit | No description |
| Pfam | PF00010 | 1.9E-11 | 403 | 449 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 8.4E-19 | 403 | 464 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 6.3E-18 | 405 | 454 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 577 aa Download sequence Send to blast |
MNMSGMGWGD EEESVVSAVL GNLASDFLRA NSNSNQNLFL VMGTDDTLNK KLSSLVDCPN 60 SENFSWNYAI FWQQTMSRSG HQVLGWGDGC CREPNEEEES KVVRVYNCNS LGLEEERWED 120 MRKRVLQKLH RLFGGSDEDN YALSLEKVTA TEIFFLASMY FFFNHGEGGP GRCYASGKHV 180 WLSDAVNSDS DYCFRSFMAK SAGIRTIVMV PTDAGVLELG SVWSLPENVD LVRSVQALFM 240 RRVKPPLMVN STNTNMSGGG IHKLFGKELN NSEQGSHHAF PRKLDVIRNV DERFTPQIWQ 300 GYNNKGSTFG YTPQRVDVKV QENVNVVVDD NNYKMQIQFA GSSVAASSNP STNTHLEKSG 360 SCTEKRPASL LLAGGAGTAS VADDKRPRKR GRKPANGREE PLNHVEAERQ RREKLNQRFY 420 ALRSVVPNIS KMDKASLLGD AISYIKELQE KVKIMEAERV RTESSLSESD QMTTRTVESP 480 EVDIQAAKEE VVVRVTSPLD SHPASRIIQA MRNSEVSLME SKLSLADDTM FHTFVIKSNN 540 GSDPLTKEKL IAAVYPQTSS MQQPLPSSSS QVSGDN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 3e-29 | 397 | 468 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_B | 3e-29 | 397 | 468 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_E | 3e-29 | 397 | 468 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_F | 3e-29 | 397 | 468 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_G | 3e-29 | 397 | 468 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_I | 3e-29 | 397 | 468 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_M | 3e-29 | 397 | 468 | 2 | 73 | Transcription factor MYC2 |
| 5gnj_N | 3e-29 | 397 | 468 | 2 | 73 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 384 | 392 | KRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator. Regulates positively abscisic acid (ABA) response. Confers drought tolerance and sensitivity to ABA. {ECO:0000269|PubMed:17828375}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00101 | PBM | Transfer from AT2G46510 | Download |
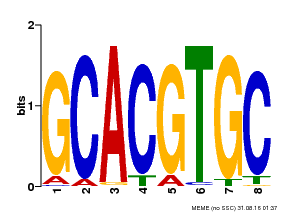
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10001374m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Transiently by ABA and polyethylene glycol (PEG). {ECO:0000269|PubMed:17828375}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006397843.1 | 0.0 | transcription factor ABA-INDUCIBLE bHLH-TYPE | ||||
| Swissprot | Q9ZPY8 | 0.0 | AIB_ARATH; Transcription factor ABA-INDUCIBLE bHLH-TYPE | ||||
| TrEMBL | V4N299 | 0.0 | V4N299_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006397843.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3758 | 27 | 59 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46510.1 | 0.0 | ABA-inducible BHLH-type transcription factor | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10001374m |
| Entrez Gene | 18015525 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



