 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10000853m | ||||||||
| Common Name | EUTSA_v10000853mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 535aa MW: 59144.1 Da PI: 5.9109 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 38.2 | 2.6e-12 | 364 | 410 | 3 | 54 |
HHHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 3 rahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
++h e+Er RRd++N++f Lr ++P+ K++Ka+ L Av YI++L
Thhalv10000853m 364 MNHVEAERLRRDKLNQRFYALRAVVPNV-----SKMDKASLLGDAVCYINEL 410
589************************6.....5***************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.1E-48 | 41 | 225 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.52 | 361 | 410 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.57E-16 | 364 | 432 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.65E-12 | 364 | 411 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 2.0E-15 | 365 | 431 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.3E-9 | 365 | 410 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.3E-14 | 367 | 416 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005509 | Molecular Function | calcium ion binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 535 aa Download sequence Send to blast |
MNHFNTDHNL SIIEDLLTSS SFSDLWPPLP PANLNLDDNT LQKRLQAVLN GTHEAWTYVI 60 FWKPSYYEFS GESVLIWGDG IHKDDDVKSR RRKKTMAEKE HRSKVLRELN SMISGEAFPV 120 VGGDDDDDDG DVEVTDTEWY FLVSMTCSFC IGSGLAGKAF ATYNPVWVTG SDDINGSGCD 180 RAKQGGGLGL ETIVCIPLDN GVLELGSTVE IRQHSDLLNK IRFLFNFKGS DDFPGAPNSK 240 SKLFSTQVAK VSLRTVTDNP NPTYPKIQKS FSPELNFSMV TPNSSTGALS GEILNFGDDV 300 KRNPGILNCT SYPGQIQNDD GDLLAAVKKI DDSDQSNINA TVVCENKRQS KRGRKPANGR 360 KEPMNHVEAE RLRRDKLNQR FYALRAVVPN VSKMDKASLL GDAVCYINEL KSRAENAEME 420 KIAIEIQLQK LKEEISGCNA ISSVCADGKN ASKIGTAKID VKVMGCDAMI RVESSKRNHP 480 GARLMNAFMD LELEVNHASV SVINDLMVQQ ATVKMVLKFY TEEQLRVMLI SKII* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4rqw_A | 4e-50 | 36 | 227 | 4 | 194 | Transcription factor MYC3 |
| 4rqw_B | 4e-50 | 36 | 227 | 4 | 194 | Transcription factor MYC3 |
| 4rs9_A | 4e-50 | 36 | 227 | 4 | 194 | Transcription factor MYC3 |
| 4yz6_A | 4e-50 | 36 | 227 | 4 | 194 | Transcription factor MYC3 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00547 | DAP | Transfer from AT5G46830 | Download |
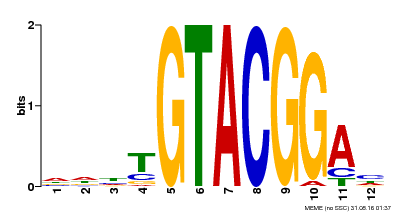
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10000853m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By UV treatment. {ECO:0000269|PubMed:12679534}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006398377.1 | 0.0 | transcription factor bHLH28 | ||||
| Swissprot | Q9LUK7 | 0.0 | BH028_ARATH; Transcription factor bHLH28 | ||||
| TrEMBL | V4KPJ8 | 0.0 | V4KPJ8_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006398377.1 | 0.0 | (Eutrema salsugineum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM13953 | 16 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G46830.1 | 0.0 | NACL-inducible gene 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10000853m |
| Entrez Gene | 18016238 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



