 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thecc1EG011484t1 | ||||||||
| Common Name | TCM_011484 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Malvales; Malvaceae; Byttnerioideae; Theobroma
|
||||||||
| Family | YABBY | ||||||||
| Protein Properties | Length: 239aa MW: 26137.1 Da PI: 8.6545 | ||||||||
| Description | YABBY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | YABBY | 59.1 | 1.9e-18 | 77 | 124 | 1 | 48 |
YABBY 1 advfssseqvCyvqCnfCntilavsvPstslfkvvtvrCGhCtsllsv 48
+d++ +se++Cyv+CnfCnt+lav +P + l+ +vtv+CGhC++l +
Thecc1EG011484t1 77 MDLVPQSEHLCYVRCNFCNTVLAVGIPCKRLLDTVTVKCGHCSNLSFL 124
578899*************************************98544 PP
| |||||||
| 2 | YABBY | 95.2 | 1.6e-29 | 161 | 222 | 102 | 163 |
YABBY 102 senedeevprvppvirPPekrqrvPsaynrfikeeiqrikasnPdishreafsaaaknWahf 163
s +++ ++p++p v++PPek+ r Psaynrf+keeiqrika+nP+i hreafsaaaknWa +
Thecc1EG011484t1 161 SPSSEPSSPKAPFVVKPPEKKHRLPSAYNRFMKEEIQRIKAANPEIPHREAFSAAAKNWARY 222
5566778999999***********************************************76 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04690 | 6.3E-52 | 82 | 222 | IPR006780 | YABBY protein |
| SuperFamily | SSF47095 | 1.44E-8 | 166 | 220 | IPR009071 | High mobility group box domain |
| Gene3D | G3DSA:1.10.30.10 | 4.8E-5 | 176 | 221 | IPR009071 | High mobility group box domain |
| CDD | cd01390 | 7.47E-6 | 183 | 219 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010254 | Biological Process | nectary development | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0048479 | Biological Process | style development | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 239 aa Download sequence Send to blast |
MPPKGPGSLV FFVHQEDNSV LEKDIKNPNL FSLLLKPTAP LFTSLAALCS ILFSSLLFPS 60 LPFPSFDNMN LEEKVGMDLV PQSEHLCYVR CNFCNTVLAV GIPCKRLLDT VTVKCGHCSN 120 LSFLSTRPPL QGQCLDPQTS LSLQSFCGDF RKGQSPSPSS SPSSEPSSPK APFVVKPPEK 180 KHRLPSAYNR FMKEEIQRIK AANPEIPHRE AFSAAAKNWA RYIPNSPATS VSGSRSNE* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor required for the initiation of nectary development. Also involved in suppressing early radial growth of the gynoecium, in promoting its later elongation and in fusion of its carpels by regulating both cell division and expansion. Establishes the polar differentiation in the carpels by specifying abaxial cell fate in the ovary wall. Regulates both cell division and expansion. {ECO:0000269|PubMed:10225997, ECO:0000269|PubMed:10225998, ECO:0000269|PubMed:10535738, ECO:0000269|PubMed:11714690, ECO:0000269|PubMed:15598802, ECO:0000269|Ref.10}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00220 | DAP | Transfer from AT1G69180 | Download |
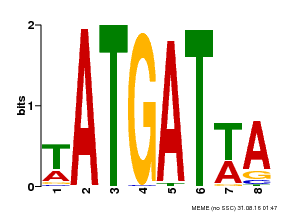
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by SPT and by A class genes AP2 and LUG in the outer whorl. In the third whorl, B class genes AP3 and PI, and the C class gene AG act redundantly with each other and in combination with SEP1, SEP2, SEP3, SHP1 and SHP2 to activate CRC in nectaries and carpels. LFY enhances its expression. {ECO:0000269|PubMed:10225998, ECO:0000269|PubMed:15598802}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY854804 | 1e-178 | AY854804.1 Gossypium hirsutum isolate 1 CRABS CLAW mRNA, partial cds. | |||
| GenBank | AY854805 | 1e-178 | AY854805.1 Gossypium hirsutum isolate 2 CRABS CLAW mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017970271.1 | 1e-121 | PREDICTED: protein CRABS CLAW isoform X2 | ||||
| Swissprot | Q8L925 | 2e-69 | CRC_ARATH; Protein CRABS CLAW | ||||
| TrEMBL | A0A061EB61 | 1e-172 | A0A061EB61_THECC; Plant-specific transcription factor YABBY family protein | ||||
| STRING | EOY01637 | 1e-173 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM9725 | 28 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69180.1 | 3e-71 | YABBY family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thecc1EG011484t1 |
| Entrez Gene | 18610215 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



