 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_135600_rufh.t1 | ||||||||
| Common Name | SOVF_135600 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 337aa MW: 36441.4 Da PI: 7.5026 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.5 | 2.7e-32 | 172 | 227 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFv+a++qLGGs++AtPk+i++ mkv+gLt+++vkSHLQkYRl+
Sp_135600_rufh.t1 172 KARRCWSPELHRRFVHALQQLGGSHVATPKQIRDSMKVDGLTNDEVKSHLQKYRLH 227
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 13.716 | 169 | 229 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-28 | 169 | 230 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 9.86E-17 | 170 | 230 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.6E-26 | 172 | 227 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.8E-7 | 174 | 225 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 337 aa Download sequence Send to blast |
CFNGHSECSE QTTSDGPILE EFIPLKRSNS PDDDDELESN KPQINNNDKS GKKSDWLRSV 60 QLWNQTPDPP TTTQEDHQPK KVAVVEVKRN GSVGGAFHPF QKEKNNNGDE SDNNNTNNKN 120 NSKKQATSTP AASACSTVQE EEMGPSSGGS GGSDGGGGGS SKKEKDAQSQ RKARRCWSPE 180 LHRRFVHALQ QLGGSHVATP KQIRDSMKVD GLTNDEVKSH LQKYRLHTRR PSSAHHNNGN 240 QQSPQFVVVG GIWVPPEYAA TTARDTTTTV AASGGIYAPI ATVAAPPRPM HQTSTQRQSS 300 DQSHCEERGS HSEGHVVHSN SPTTSSSTHT TTASHAQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 1e-13 | 172 | 225 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-13 | 172 | 225 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-13 | 172 | 225 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-13 | 172 | 225 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
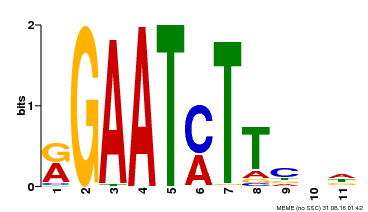
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021857301.1 | 0.0 | transcription factor HHO2-like isoform X1 | ||||
| Swissprot | Q9FPE8 | 3e-76 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A0K9QVW3 | 0.0 | A0A0K9QVW3_SPIOL; Uncharacterized protein (Fragment) | ||||
| STRING | XP_010695844.1 | 1e-157 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68670.1 | 5e-64 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



