 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_121600_fyyw.t1 | ||||||||
| Common Name | SOVF_121600 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 327aa MW: 35566.5 Da PI: 8.7721 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 24.3 | 7.4e-08 | 6 | 52 | 3 | 45 |
SS-HHHHHHHHHHHHHTTTT.....-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg.....tWktIartmgkgRtlkqcksrwq 45
+W+ eE++ + +a++ + + +W+++a+ + ++t +++k ++
Sp_121600_fyyw.t1 6 SWSREEEKAFENAIATYSIDqsnedDWDKVAAVVT-TKTIEEIKQHYE 52
6*********************************9.**********95 PP
| |||||||
| 2 | Myb_DNA-binding | 40.3 | 7.2e-13 | 145 | 189 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + ++G+g+W++I+r + +Rt+ q+ s+ qky
Sp_121600_fyyw.t1 145 PWTEEEHRLFLLGLDKFGKGDWRSISRNFVITRTPTQVASHAQKY 189
7*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 9.82 | 1 | 59 | IPR017884 | SANT domain |
| SMART | SM00717 | 1.2E-6 | 3 | 57 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.29E-10 | 6 | 61 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 2.5E-6 | 6 | 52 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 6.05E-8 | 6 | 55 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 3.8E-4 | 6 | 57 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 17.466 | 138 | 194 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.47E-17 | 139 | 195 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.1E-17 | 141 | 192 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.3E-10 | 142 | 192 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.3E-11 | 143 | 188 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 5.88E-10 | 145 | 190 | No hit | No description |
| Pfam | PF00249 | 1.9E-11 | 145 | 189 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 327 aa Download sequence Send to blast |
MPNLASWSRE EEKAFENAIA TYSIDQSNED DWDKVAAVVT TKTIEEIKQH YELLVEDVGA 60 IEAGKVPLPK YASEETNHHN NSSNNNGTLL STSTTSKDDH SPSFSKEQKG NSNQGNGYSA 120 FGPSSMGNST KASSKLEQER KKGIPWTEEE HRLFLLGLDK FGKGDWRSIS RNFVITRTPT 180 QVASHAQKYF IRLNSMNRDR RRSSIHDITS VNGGGLSPNQ VPITGQQNNN PAAPAVGTAA 240 AAAAANKHRA PPNVNVNMAA AGIGMYGTQM GHPISAPPPP PHHMVSAVGT PVMLPPPGHP 300 HHHHHHHHPP HYAVPVAYPV PPPPMHR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 4e-15 | 7 | 73 | 11 | 75 | RADIALIS |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00191 | DAP | Transfer from AT1G49010 | Download |
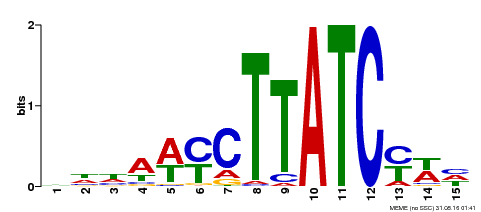
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021854462.1 | 0.0 | transcription factor MYBS1-like | ||||
| TrEMBL | A0A0K9R1X8 | 0.0 | A0A0K9R1X8_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010673224.1 | 1e-164 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49010.1 | 2e-64 | MYB family protein | ||||



