 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_117570_tude.t1 | ||||||||
| Common Name | SOVF_117570 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 436aa MW: 48819.2 Da PI: 7.4396 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.5 | 5.3e-32 | 98 | 151 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH +Fv+ave+LGG+e+AtPk +l++m++kgL+++hvkSHLQ+YR+
Sp_117570_tude.t1 98 PRLRWTPDLHLCFVHAVERLGGQERATPKLVLQMMNIKGLSIAHVKSHLQMYRS 151
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 5.5E-30 | 94 | 152 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.957 | 94 | 154 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.69E-14 | 96 | 152 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-22 | 98 | 153 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.1E-9 | 99 | 150 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 436 aa Download sequence Send to blast |
MIRAGSPNTQ GSKTSLSNNH KVDEESEEDD KVLLLEEEEE EEEEEEVGEE VDDNSNSNSR 60 VMIKINGSSS STSTVEEDDK KGPNSSGSVR PYNRSKMPRL RWTPDLHLCF VHAVERLGGQ 120 ERATPKLVLQ MMNIKGLSIA HVKSHLQMYR SKKIDESNQA AMSNQGIFSD NGRDNIYNLS 180 QLPMLQGYNH QRPPFSLRYG DPSWRGHSNQ AYAPYNVGTS SSERSRNGLN FSSLPERMIF 240 SSSNSNIFQG QMSNWRNQQT LKKTPIILSH QSNGSSSCQS TSRPIRSNME LNLAISPIQQ 300 VVPRDQGKDH QRSNSSNFLK PYSNIIDIKQ GSTTTTSNHL KRKAMDLDLS LALAHPKDID 360 NNDPLTKKNT KDKEVIIGSS DKLLSLSLQY PPPPLLLSSS SEHDNQFIIS KLKEGDASSS 420 RMNNKQQTRT SLDLTL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 6e-18 | 99 | 153 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 6e-18 | 99 | 153 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 6e-18 | 99 | 153 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 6e-18 | 99 | 153 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 6e-18 | 99 | 153 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 6e-18 | 99 | 153 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 6e-18 | 99 | 153 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 6e-18 | 99 | 153 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 6e-18 | 99 | 153 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 6e-18 | 99 | 153 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 6e-18 | 99 | 153 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 6e-18 | 99 | 153 | 4 | 58 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
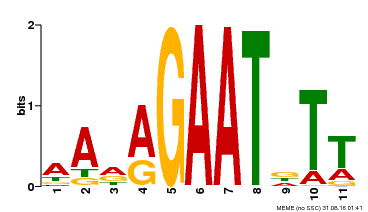
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00297 | DAP | Transfer from AT2G38300 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021841563.1 | 0.0 | probable transcription factor KAN2 isoform X1 | ||||
| TrEMBL | A0A0K9R1I5 | 0.0 | A0A0K9R1I5_SPIOL; Uncharacterized protein | ||||
| STRING | XP_010693076.1 | 1e-154 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38300.1 | 6e-53 | G2-like family protein | ||||



