 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sp_046550_watp.t1 | ||||||||
| Common Name | SOVF_046550 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Chenopodiaceae; Chenopodioideae; Anserineae; Spinacia
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 507aa MW: 55801.2 Da PI: 5.2537 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50.6 | 4.5e-16 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT eEd++lv ++ +G +W++ ++ g+ R++k+c++rw +yl
Sp_046550_watp.t1 14 KGPWTSEEDQILVTFIHNHGHSNWRALPKQAGLLRCGKSCRLRWINYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 48.4 | 2.1e-15 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg+++ eE++ +++++++lG++ W++Ia++++ gRt++++k+ w+++l
Sp_046550_watp.t1 67 RGNFSREEEDSIIQLHEMLGNR-WSAIAARLP-GRTDNEIKNVWHTHL 112
89********************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.5E-23 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.912 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.11E-29 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.0E-12 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.5E-14 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.09E-10 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 24.629 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-25 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.0E-14 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.6E-14 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.07E-10 | 69 | 112 | No hit | No description |
| PROSITE profile | PS50873 | 53.731 | 240 | 495 | IPR002016 | Haem peroxidase, plant/fungal/bacterial |
| Gene3D | G3DSA:1.10.520.10 | 1.7E-22 | 254 | 348 | No hit | No description |
| Pfam | PF00141 | 1.2E-53 | 254 | 462 | IPR002016 | Haem peroxidase, plant/fungal/bacterial |
| SuperFamily | SSF48113 | 8.25E-67 | 260 | 495 | IPR010255 | Haem peroxidase |
| CDD | cd00693 | 9.76E-115 | 262 | 494 | No hit | No description |
| PRINTS | PR00461 | 1.2E-22 | 272 | 285 | IPR000823 | Plant peroxidase |
| PRINTS | PR00461 | 1.2E-22 | 291 | 301 | IPR000823 | Plant peroxidase |
| PRINTS | PR00458 | 1.6E-16 | 292 | 309 | IPR002016 | Haem peroxidase, plant/fungal/bacterial |
| PRINTS | PR00461 | 1.2E-22 | 310 | 325 | IPR000823 | Plant peroxidase |
| PRINTS | PR00458 | 1.6E-16 | 310 | 322 | IPR002016 | Haem peroxidase, plant/fungal/bacterial |
| Gene3D | G3DSA:1.10.420.10 | 6.8E-37 | 349 | 477 | No hit | No description |
| PRINTS | PR00461 | 1.2E-22 | 356 | 368 | IPR000823 | Plant peroxidase |
| PRINTS | PR00458 | 1.6E-16 | 357 | 372 | IPR002016 | Haem peroxidase, plant/fungal/bacterial |
| PRINTS | PR00461 | 1.2E-22 | 416 | 431 | IPR000823 | Plant peroxidase |
| PRINTS | PR00458 | 1.6E-16 | 418 | 433 | IPR002016 | Haem peroxidase, plant/fungal/bacterial |
| PRINTS | PR00461 | 1.2E-22 | 432 | 449 | IPR000823 | Plant peroxidase |
| PRINTS | PR00461 | 1.2E-22 | 472 | 485 | IPR000823 | Plant peroxidase |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006979 | Biological Process | response to oxidative stress | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0055114 | Biological Process | oxidation-reduction process | ||||
| GO:0098869 | Biological Process | cellular oxidant detoxification | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0004601 | Molecular Function | peroxidase activity | ||||
| GO:0020037 | Molecular Function | heme binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 507 aa Download sequence Send to blast |
MGRAPCCDKV GLKKGPWTSE EDQILVTFIH NHGHSNWRAL PKQAGLLRCG KSCRLRWINY 60 LKPDIKRGNF SREEEDSIIQ LHEMLGNRWS AIAARLPGRT DNEIKNVWHT HLKKRLKEYE 120 PTKSSSRGKT SKKSRANNNN AAVPTTTTDS NTSGGNDSFP ESSSIATSGS PQQSSTTTTT 180 TTSDDYFSSG TEVSSSVELF PNSDTSLFGS PSLEYLPIID ETFWESSSSQ EVALDTSEFD 240 YIVEDNDQIV QFESFLREED QEKEAGANAS VREYELIDEI KNFLEQACRG IVSCADIITL 300 ATRDAIALAG GPVYNVPTGR LDGLISRASE VNLPGPASTV AQIRQTFRAR GMTVRDMVLL 360 VGGGHSIGVA HCNFFQDRLF NFQGTGAPDS TMDTTLLNEL QGLCGFSSQV PNPTTFLDQN 420 TSFVLDNEIY KQMKMNRGIL QIDQELALDP LSSKIVSDFA ADNGFFRRRF VDAIIKMGNL 480 PGSGGEVRGN CRRINSATAQ QSLLSSI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1pa2_A | 2e-50 | 262 | 495 | 65 | 304 | PEROXIDASE |
| 1qo4_A | 2e-50 | 262 | 495 | 65 | 304 | PEROXIDASE |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00375 | DAP | Transfer from AT3G23250 | Download |
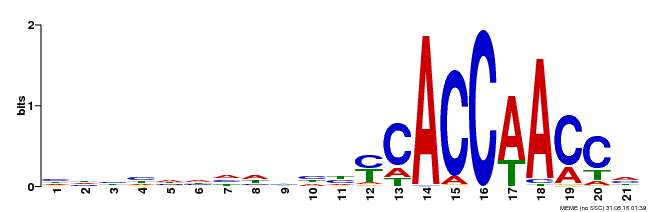
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021847645.1 | 0.0 | transcription factor MYB30-like | ||||
| TrEMBL | A0A0K9RNI0 | 0.0 | A0A0K9RNI0_SPIOL; Peroxidase | ||||
| STRING | XP_010695531.1 | 1e-112 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G23250.1 | 7e-79 | myb domain protein 15 | ||||



