 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.007G155900.5.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 350aa MW: 36174.7 Da PI: 5.2822 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 47.7 | 3.5e-15 | 286 | 337 | 5 | 56 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkev 56
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L+++N++L+k+ ee+ ++
Sobic.007G155900.5.p 286 RRQRRMIKNRESAARSRARKQAYTMELEAEVQKLKEQNEELQKKQEEIMEMQ 337
79******************************************99999986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 6.4E-14 | 282 | 347 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.484 | 284 | 335 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 2.7E-13 | 286 | 338 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 4.0E-11 | 286 | 335 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 7.4E-15 | 286 | 335 | No hit | No description |
| CDD | cd14707 | 2.44E-27 | 286 | 340 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 289 | 304 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0010255 | Biological Process | glucose mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 350 aa Download sequence Send to blast |
MDPKDGERMG AAGPGPLSRQ GSIYSLTFDE FQNTLGGMGG GLGKDFGSMN MDELLRSIWT 60 AEESQAIASA SASASASAAG AGAGPVGDGG AALQRQGSLT LPRTLSVKTV DEVWRDFARE 120 GPPGPTAGGA EPQPNRQPTL GEMTLEEFLV RAGVVRDNPA AAAAAAAAAV SAQPVAPRPI 180 QAVNNGASIF FGNFGGANDA GAGAMGFAPV GIGDQAMGNG LMPGVAGMAG GAVTVVSPVD 240 TSVAQLDSMG KGNGDLSSPM APVPYPFEGV IRGRRSGAGV EKVVERRQRR MIKNRESAAR 300 SRARKQAYTM ELEAEVQKLK EQNEELQKKQ EEIMEMQKNQ VYQKLHNVC* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Sbi.21427 | 4e-34 | shoot | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, leaves and embryos. {ECO:0000269|PubMed:10611387}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that mediates abscisic acid (ABA) signaling. Binds specifically to the ABA-responsive element (ABRE) of the EMP1 and RAB16A gene promoters. {ECO:0000269|PubMed:10611387}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00186 | DAP | Transfer from AT1G45249 | Download |
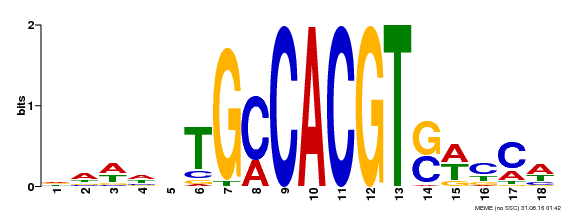
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.007G155900.5.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA). {ECO:0000269|PubMed:10611387}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT039652 | 0.0 | BT039652.1 Zea mays full-length cDNA clone ZM_BFc0037E11 mRNA, complete cds. | |||
| GenBank | EU964768 | 0.0 | EU964768.1 Zea mays clone 281458 bZIP transcription factor ABI5 mRNA, complete cds. | |||
| GenBank | KJ726943 | 0.0 | KJ726943.1 Zea mays clone pUT3489 bZIP transcription factor (bZIP68) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021320779.1 | 0.0 | bZIP transcription factor TRAB1-like isoform X1 | ||||
| Swissprot | Q6ZDF3 | 1e-136 | TRAB1_ORYSJ; bZIP transcription factor TRAB1 | ||||
| TrEMBL | A0A1Z5RA36 | 0.0 | A0A1Z5RA36_SORBI; Uncharacterized protein | ||||
| STRING | Si013904m | 1e-174 | (Setaria italica) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G19290.2 | 9e-34 | ABRE binding factor 4 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.007G155900.5.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



