 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.004G281800.1.p | ||||||||
| Common Name | Sb04g031820, SORBIDRAFT_04g031820 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 282aa MW: 30879.7 Da PI: 9.508 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 42.8 | 1.3e-13 | 36 | 80 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT++E++++++a +++ + Wk+I + +g +t q++s+ qky
Sobic.004G281800.1.p 36 RESWTEQEHDKFLEALQLFDRD-WKKIEAFVG-SKTVIQIRSHAQKY 80
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 8.22E-17 | 30 | 86 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 15.408 | 31 | 85 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.2E-18 | 34 | 83 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 8.8E-7 | 34 | 86 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.3E-11 | 35 | 83 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.1E-11 | 36 | 80 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.39E-8 | 38 | 81 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 282 aa Download sequence Send to blast |
MVSASAPPPP LQSDAAGSGE DASKKVRKPY TITKSRESWT EQEHDKFLEA LQLFDRDWKK 60 IEAFVGSKTV IQIRSHAQKY FLKVQKNGTS EHVPPPRPKR KAAHPYPQKA SKNEPNYGLK 120 TDSSSIHRNS GMNVSVSSWA HSSIPQAVAS TMVKDLGAGT PGPNNFCSSS TEGPPRTWQP 180 GETNDQINQV PSLRLMPDFA EVYSFLGSVF DPSTSGHLQK LKEMNPIDVE TALLLMRNLS 240 INLTSPDFED QRKLLSSYST SDGLQLGSSR SSALAMSTPF M* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00625 | PBM | Transfer from PK02532.1 | Download |
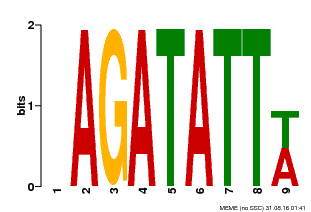
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.004G281800.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HF679446 | 0.0 | HF679446.1 Saccharum hybrid cultivar Co 86032 mRNA for ScMYB40 protein. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021313806.1 | 0.0 | protein REVEILLE 6 isoform X4 | ||||
| Swissprot | Q8H0W3 | 1e-100 | RVE6_ARATH; Protein REVEILLE 6 | ||||
| TrEMBL | A0A194YRZ7 | 0.0 | A0A194YRZ7_SORBI; Uncharacterized protein | ||||
| STRING | Sb04g031820.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1096 | 38 | 111 | Representative plant | OGRP928 | 15 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52660.2 | 9e-89 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.004G281800.1.p |
| Entrez Gene | 8076033 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



