 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.003G004500.1.p | ||||||||
| Common Name | Sb03g000570, SORBIDRAFT_03g000570 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 623aa MW: 67721.5 Da PI: 7.6102 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.3 | 1.2e-12 | 468 | 514 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
Sobic.003G004500.1.p 468 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAITYITDLQ 514
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 1.6E-50 | 62 | 247 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.831 | 464 | 513 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 5.76E-19 | 467 | 531 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 4.86E-15 | 467 | 518 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 2.3E-18 | 468 | 527 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.8E-10 | 468 | 514 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.9E-16 | 470 | 519 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 623 aa Download sequence Send to blast |
MVMKMELEDD GAMGGRGTGG MWTEEDRALG AAVLGADAFA YLTKGGGAIS EGLVAASLPD 60 DLQNKLQELV ESESPGTGWN YAIFWQLSRT KSGDLVLGWG DGSCREPRDG EVGAAASAGS 120 DDTKQRMRKR VLQRLHIAFG VADEEDYAPG IDQVTDTEMF FLASMYFAFP RRTGGPGQAF 180 AAGIPLWVPN SERKVFPANY CYRGFLANAA GFRTIVLVPF ESGVLELGSM QHIAESSDTI 240 QSIRSVFAGT SGNKTAVQRH EGNGPAPPER SPSLAKIFGK DLNLGRPSVG PAVGVSKVDE 300 RTWEQRSAAG GTSLLPNVQK GLQNFTWSQA RGLNSHQQKF GNGVLIVSNE AAHRNNGAVD 360 SPSAAQFQLQ KAPQLQKLQL QKLPVVQKTP QLVNQQPMQP QVPRQIDFSA GSSSKPGVLV 420 TRAAAVLDGE SAEVDGLCKE EGPPPVIEDR RPRKRGRKPA NGREEPLNHV EAERQRREKL 480 NQRFYALRAV VPNISKMDKA SLLGDAITYI TDLQKKLKEM ESERERLLES GMVDPRERAP 540 RPEVDIQVVQ DEVLVRVMSP MDNHPVRKVF QAFEEAEVRV GESKVTGNNN GTVVHSFIIK 600 CPGAEQQTRE KVIAAMSRAM SS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 3e-28 | 462 | 524 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_B | 3e-28 | 462 | 524 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_E | 3e-28 | 462 | 524 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_F | 3e-28 | 462 | 524 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_G | 3e-28 | 462 | 524 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_I | 3e-28 | 462 | 524 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_M | 3e-28 | 462 | 524 | 2 | 64 | Transcription factor MYC2 |
| 5gnj_N | 3e-28 | 462 | 524 | 2 | 64 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 449 | 457 | RRPRKRGRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
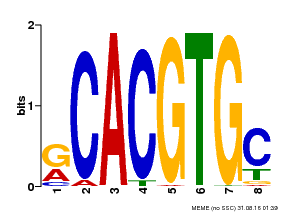
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.003G004500.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT054953 | 0.0 | BT054953.1 Zea mays full-length cDNA clone ZM_BFc0103E04 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021312143.1 | 0.0 | transcription factor bHLH13 isoform X1 | ||||
| TrEMBL | C5XJK2 | 0.0 | C5XJK2_SORBI; Uncharacterized protein | ||||
| STRING | Sb03g000570.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5174 | 38 | 60 | Representative plant | OGRP2531 | 11 | 23 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 8e-72 | bHLH family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.003G004500.1.p |
| Entrez Gene | 8078088 |



