 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sobic.001G077800.1.p | ||||||||
| Common Name | Sb01g007030, SORBIDRAFT_01g007030 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Andropogonodae; Andropogoneae; Sorghinae; Sorghum
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 383aa MW: 38888.3 Da PI: 7.8296 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.7 | 2.4e-32 | 117 | 170 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
pr+rWt+ LH+rFv+ave LGG+e+AtPk+++elm+vk+Ltl+hvkSHLQ+YR+
Sobic.001G077800.1.p 117 PRMRWTTALHARFVHAVELLGGHERATPKSVMELMNVKDLTLAHVKSHLQMYRT 170
9****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 9.85E-16 | 115 | 171 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 4.6E-29 | 115 | 171 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 3.2E-23 | 117 | 170 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.7E-8 | 118 | 169 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005618 | Cellular Component | cell wall | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 383 aa Download sequence Send to blast |
MLQRNLPNLC LQISPPAASS AVVASDAPVA ASAKPPPTQP ISTATYAEGS GEVGFFANPS 60 PGAEPPGLSL GLATPALGDN AAGRRGHHLQ PQGCAFKRAA ARASLPAAGS KRSARAPRMR 120 WTTALHARFV HAVELLGGHE RATPKSVMEL MNVKDLTLAH VKSHLQMYRT VKSTDRSLHT 180 ATGEALPLQR TAATGMEAAA AAAAGGGGGG AGGGVIMVPV PAACDDMVGI CSSPSAGSAP 240 PAMATTSAAH FLCAPAATTA PLAAAPSPPP PIPPRSRTTD HAAAVPEKGV AIVDSLHRPV 300 LEDTHQAAQE KANNGQLPMG LHGCVDLLDP MATNCSSPAS SSPSLASLEQ LTADMYAPNL 360 EISLGRQDWS MDRPEGLCLK YL* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 1e-16 | 118 | 172 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-16 | 118 | 172 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-16 | 118 | 172 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-16 | 118 | 172 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Sbi.21091 | 0.0 | panicle | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00107 | PBM | Transfer from AT5G42630 | Download |
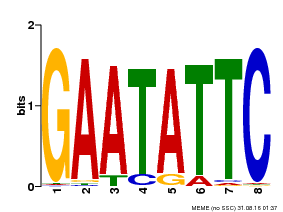
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sobic.001G077800.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT065124 | 1e-169 | BT065124.1 Zea mays full-length cDNA clone ZM_BFb0205B22 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002466395.1 | 0.0 | probable transcription factor GLK2 | ||||
| TrEMBL | C5WZ98 | 0.0 | C5WZ98_SORBI; Uncharacterized protein | ||||
| STRING | Sb01g007030.1 | 0.0 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6554 | 37 | 50 | Representative plant | OGRP570 | 15 | 80 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42630.2 | 6e-37 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sobic.001G077800.1.p |
| Entrez Gene | 8086186 |



