 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sme2.5_03132.1_g00006.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Solanoideae; Solaneae; Solanum
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 826aa MW: 90131.2 Da PI: 6.3162 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.6 | 6.8e-21 | 131 | 186 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l L++rqVk+WFqNrR+++k
Sme2.5_03132.1_g00006.1 131 KKRYHRHTPQQIQELESLFKECPHPDEKQRLELSKRLCLETRQVKFWFQNRRTQMK 186
688999***********************************************999 PP
| |||||||
| 2 | START | 202.2 | 2.1e-63 | 338 | 562 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla 77
ela++a++elvk+a+ +ep+W++s e++n +e++++f++ + + +ea+r++g+v+ ++ lve+l+d++ +W e+++
Sme2.5_03132.1_g00006.1 338 ELALAAMDELVKMAQTDEPLWFRSIegvrEILNHEEYIRTFTPCIGmkpnsFVSEASRETGMVIINSLALVETLMDSN-KWAEMFP 422
5899**************************************99989***9***************************.******* PP
....EEEEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEE CS
START 78 ....kaetlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgi 152
+ +t++vissg galqlm aelq+lsplvp R++ f+R+++q+ +g+w++vdvS+d ++ + + + +++lpSg+
Sme2.5_03132.1_g00006.1 423 cliaRTSTTDVISSGmggtrnGALQLMHAELQVLSPLVPiREVNFLRFCKQHAEGVWAVVDVSIDTIRETSGAPTFPNCRRLPSGC 508
*********************************************************************999************** PP
EEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 153 liepksnghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqcek 206
++++++ng+skvtwveh+++++ h l+r+l++ g+ +ga++wvatlqrqce+
Sme2.5_03132.1_g00006.1 509 VVQDMPNGYSKVTWVEHAEYDEGANHHLYRQLISAGMGFGAQRWVATLQRQCEC 562
****************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.01E-20 | 118 | 188 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-21 | 118 | 188 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.248 | 128 | 188 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.2E-17 | 129 | 192 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.88E-18 | 130 | 188 | No hit | No description |
| Pfam | PF00046 | 2.0E-18 | 131 | 186 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 163 | 186 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 43.397 | 329 | 565 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.66E-31 | 331 | 562 | No hit | No description |
| CDD | cd08875 | 7.95E-123 | 333 | 561 | No hit | No description |
| Pfam | PF01852 | 7.6E-55 | 338 | 562 | IPR002913 | START domain |
| SMART | SM00234 | 1.3E-48 | 338 | 562 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.1E-23 | 581 | 812 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 826 aa Download sequence Send to blast |
MNIGGFLDNN SGGGGARIVA DIPFNNNANS SSNSNNKNNN NSSMPTGAIS QPRLVPQSLA 60 KTMFNSAGLS LALQTGMEGQ GEVTRMGENY EGNNSVGRRS REEEPDSRSG SDNLEGASGD 120 EQDAADKPPR KKRYHRHTPQ QIQELESLFK ECPHPDEKQR LELSKRLCLE TRQVKFWFQN 180 RRTQMKTQLE RHENSILRQE NDKLRAENMS IREAMRNPIC TNCGGPAMIG EISLEEQHLR 240 IENARLKDEL DRVCALAGKF LGRPISSLVT SMPPPMPNSS LELGVGSNGF GGMSNVPTTL 300 PLAPPDFGVG ISNSLPVVPS TRQSTGIERS LERSMYLELA LAAMDELVKM AQTDEPLWFR 360 SIEGVREILN HEEYIRTFTP CIGMKPNSFV SEASRETGMV IINSLALVET LMDSNKWAEM 420 FPCLIARTST TDVISSGMGG TRNGALQLMH AELQVLSPLV PIREVNFLRF CKQHAEGVWA 480 VVDVSIDTIR ETSGAPTFPN CRRLPSGCVV QDMPNGYSKV TWVEHAEYDE GANHHLYRQL 540 ISAGMGFGAQ RWVATLQRQC ECLAILMSST VSSRDHTAIT PSGRRSMLKL AQRMTNNFCA 600 GVCASTVHKW NKLCAGNVDE DVRVMTRKSV DDPGEPAGIV LSAATSVWLP VSPQRLFDFL 660 RDERLRSEWD ILSNGGPMQE MAHIAKGQDH GNCVSLLRAS AMNANQSSML ILQETCIDAA 720 GALVVYAPVD IPAMHVVMNG GDSAYVALLP SGFSIVPDGP GSRGSNGPSC NGGPDQRISG 780 SLLTVAFQIL VNSLPTAKLT VESVETVNNL ISCTVQKIKA ALQCES |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
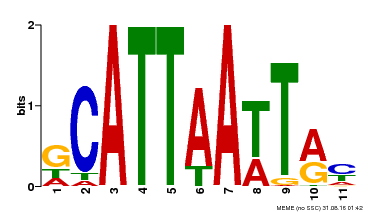
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GQ222185 | 0.0 | GQ222185.1 Solanum lycopersicum cultivar M82 cutin deficient 2 (CD2) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001234657.2 | 0.0 | cutin deficient 2 | ||||
| Refseq | XP_015059298.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A3Q7EKE6 | 0.0 | A0A3Q7EKE6_SOLLC; Uncharacterized protein | ||||
| STRING | Solyc01g091630.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1626 | 24 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



