 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sevir.9G073300.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 343aa MW: 37864.5 Da PI: 6.5125 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 29.8 | 1e-09 | 256 | 291 | 20 | 55 |
HHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 20 eknrypsaeereeLAkklgLterqVkvWFqNrRake 55
+k +yps++e+ LA+ +gL+++q+ +WF N+R ++
Sevir.9G073300.1.p 256 YKWPYPSETEKMALAETTGLDQKQINNWFINQRKRH 291
4679*****************************885 PP
| |||||||
| 2 | ELK | 41.6 | 2.7e-14 | 211 | 231 | 1 | 21 |
ELK 1 ELKhqLlrKYsgyLgsLkqEF 21
ELKhqLlrKY+gyLg+L+qEF
Sevir.9G073300.1.p 211 ELKHQLLRKYGGYLGGLRQEF 231
9*******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 1.9E-20 | 79 | 123 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 2.0E-21 | 80 | 121 | IPR005540 | KNOX1 |
| SMART | SM01256 | 1.9E-28 | 131 | 182 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 1.6E-23 | 136 | 181 | IPR005541 | KNOX2 |
| Pfam | PF03789 | 3.0E-10 | 211 | 231 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.575 | 211 | 231 | IPR005539 | ELK domain |
| SMART | SM01188 | 2.0E-7 | 211 | 232 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.989 | 231 | 294 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 7.3E-12 | 233 | 298 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 1.58E-19 | 233 | 303 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-27 | 236 | 297 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 2.43E-12 | 243 | 294 | No hit | No description |
| Pfam | PF05920 | 9.5E-17 | 251 | 290 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 269 | 292 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 343 aa Download sequence Send to blast |
MDSFRDLGGG GSSASKASFL QLPLLPASSS AQGFPSPDGH HHSSRLALQQ LLADPSAAQH 60 SHRKDGSLAQ GEISPVDAET IKTKIMSHPQ YSALVAAYLD CQKVGAPPDV SDRLSAMAAK 120 LDAQPGPSRR RHEPTRADPE LDQFMEAYCN MLVKYQEELA RPIQEAAEFF KSVERQLDSI 180 TDSNCEGAGS SEDDQDASCP EEIDPCAEDK ELKHQLLRKY GGYLGGLRQE FTKRKKKGKL 240 PKEARQKLLH WWELHYKWPY PSETEKMALA ETTGLDQKQI NNWFINQRKR HWKPTSEDMP 300 FAMMEAAAAA GGFHAPQGAA ALYMADSRTA FMAPDGMYRL GS* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during embryogenesis. {ECO:0000269|PubMed:10080693}. | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during embryogenesis. {ECO:0000269|PubMed:10080693, ECO:0000269|PubMed:9869405}. | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during embryogenesis. {ECO:0000269|PubMed:10488233}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00671 | PBM | Transfer from GRMZM2G135447 | Download |
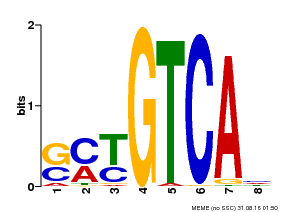
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sevir.9G073300.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT036681 | 0.0 | BT036681.1 Zea mays full-length cDNA clone ZM_BFb0131G23 mRNA, complete cds. | |||
| GenBank | EU966373 | 0.0 | EU966373.1 Zea mays clone 293957 homeobox protein rough sheath 1 mRNA, complete cds. | |||
| GenBank | JX469931 | 0.0 | JX469931.1 Zea mays subsp. mays clone UT3189 HB-type transcription factor mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004981606.1 | 0.0 | homeobox protein knotted-1-like 12 | ||||
| Swissprot | O65034 | 1e-118 | KNOSC_ORYSI; Homeobox protein knotted-1-like 12 | ||||
| Swissprot | O80416 | 1e-118 | KNOSC_ORYSJ; Homeobox protein knotted-1-like 12 | ||||
| Swissprot | Q10ED2 | 1e-119 | KNOS8_ORYSJ; Homeobox protein knotted-1-like 8 | ||||
| TrEMBL | A0A368SE34 | 0.0 | A0A368SE34_SETIT; Uncharacterized protein | ||||
| STRING | Pavir.Ia00428.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4061 | 38 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 3e-80 | KNOTTED-like from Arabidopsis thaliana | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sevir.9G073300.1.p |



