 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sevir.3G311800.2.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1394aa MW: 154879 Da PI: 7.9121 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.4 | 0.00023 | 1274 | 1297 | 1 | 22 |
EEET..TTTEEESSHHHHHHHHHH CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt 22
++C C++sFs++ +L H r
Sevir.3G311800.2.p 1274 FSCDieGCDMSFSTQQDLALHKRD 1297
899999***************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF51197 | 1.32E-24 | 95 | 334 | No hit | No description |
| SMART | SM00558 | 3.8E-46 | 168 | 337 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 35.369 | 168 | 337 | IPR003347 | JmjC domain |
| Pfam | PF02373 | 3.8E-36 | 201 | 320 | IPR003347 | JmjC domain |
| SMART | SM00355 | 9.7 | 1274 | 1296 | IPR015880 | Zinc finger, C2H2-like |
| SMART | SM00355 | 15 | 1297 | 1321 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.887 | 1297 | 1326 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1299 | 1321 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.55E-9 | 1313 | 1355 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-9 | 1324 | 1351 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.002 | 1327 | 1351 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.344 | 1327 | 1356 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1329 | 1351 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.2E-8 | 1345 | 1379 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.9E-9 | 1352 | 1380 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.284 | 1357 | 1388 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.1 | 1357 | 1383 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1359 | 1383 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1394 aa Download sequence Send to blast |
MSPPAVETPE WLRNLPVAPE YHPTAAEFVD PIAYILKIEA TVERLKASFA ANAAAAAGVD 60 GAAPAPTFPT RLQQVGFSTK NRRPASRRVW ESGERYTLEA FRAKARDIEL PRHAVPPKHA 120 TQLQLEALFW GACAARPFNV EYGNDMPGSG FAAPKEGGGG NAALAARDVG ETEWNMRLAP 180 RARGSLLRAM GRDVAGVTTP MLYVAMLYSW FAWHVEDHEL HSLNYLHFGK PKTWYGVPRD 240 AMLAFEDAVR VHGYADDLNA IMAFQTLNEK TTVLSPEVLL SAGVPCCRLV QNPGEFIITF 300 PGAYHSGFSH GFNCGEATNI ATPRWLQVAK EAAIRRASTN CGPLVSHYQL LYELALSLRP 360 RELKDSHGVP RSSRLRDKKK NEGEIMIKET FVGSVIENNN FLSILLDKSS CVIIPEIKFP 420 LPSFPTSMAP EVTVKQGLIA GTCSISQKKA EDMLAYGNAI DEIKGFEKMS ESQSSSATTS 480 SACNGRKLYE TKFGMVNSSA LLLNSEIQSG VIEKGGSHQG CGLLDQGRLP CVQCGILSYA 540 CVAIIQPKEA AVQYVISQEC MSSSAKHGEI MKSNDTSNWI TTVPPQGHSS ETDDNRIHNM 600 SSARVSDRCR QLYTSSTHGC ASALGLLASA YESSDSDEEA EAPDNISNNS ANNDAVNGIT 660 NIQSSGTSVQ YQNTNLHLYE EGCDSRATVS PMKPVENMSI AMTQASIETD MTHLADLGES 720 LTAYDQWSTY LDLDDDPTAP GVKASLSTSF SRAKGAMEPD ALTLLKYSKD SCRMHVFCLE 780 HALETWTRLQ QFGGANIVLL CHPEYPKAES AAKVIAEELG MKHAWKDITF KNATDEDIGR 840 IRLALQDEDA EPTSSDWAVK MGINIYYSAK QSKSPLYSKQ VPYNSIIYKA FAQENPDSLT 900 DERQRSGTTK KKVAGSWCGK IWMSNQVHPL LDREREEQDH DMVYSKAMVS VNSYDRIQEP 960 SARSTSLINR SLSKRVSRRK EGDSVERAKK KRYTASDVAT FDQPRNFDDH GKHESDGDEN 1020 ENEDAQKTLQ HQHESEKTNK NSSSKRRKDD KRNNNFYERH NDSDDHDYRL GIDSTENTTV 1080 GDCDNSPPQG LDVVKVKSDT QLQGSKKKSS KCKASDDLSN VEKRLQKMVK KVSTKKHKND 1140 KTNRQFQEKH SKDNNVDLLH EDNGDEATQE NWDGVQQKTN DVKVKSRGKM HSGKKKASKC 1200 QTSDGLHNGD NEAKFSCDTD VCHRDKATID KWEEIPKEKA DDVKVKSKMQ SGKKKASKHP 1260 ASDGLRNGDK GAKFSCDIEG CDMSFSTQQD LALHKRDICP VKGCKKKFFC HKYLLQHRKV 1320 HLDERPLMCS FTGCKKTFKW PWARTEHMRV HTGVRPYACT EPGCTQTFRF VSDFSRHKRK 1380 TGHSCDKKKK NST* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 1e-60 | 10 | 358 | 7 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 1e-60 | 10 | 358 | 7 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 977 | 991 | RRKEGDSVERAKKKR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
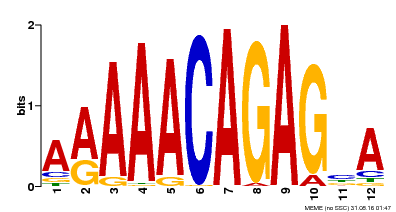
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sevir.3G311800.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025807104.1 | 0.0 | lysine-specific demethylase JMJ705-like | ||||
| TrEMBL | A0A2T7EFS1 | 0.0 | A0A2T7EFS1_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Cb01041.1.p | 0.0 | (Panicum virgatum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 1e-149 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sevir.3G311800.2.p |



