 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sevir.3G311800.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1415aa MW: 157126 Da PI: 7.9538 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 13.4 | 0.00024 | 1295 | 1318 | 1 | 22 |
EEET..TTTEEESSHHHHHHHHHH CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt 22
++C C++sFs++ +L H r
Sevir.3G311800.1.p 1295 FSCDieGCDMSFSTQQDLALHKRD 1318
899999***************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 2.7E-17 | 17 | 58 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.883 | 18 | 59 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 1.3E-13 | 19 | 52 | IPR003349 | JmjN domain |
| SuperFamily | SSF51197 | 1.37E-24 | 116 | 355 | No hit | No description |
| SMART | SM00558 | 3.8E-46 | 189 | 358 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 35.369 | 189 | 358 | IPR003347 | JmjC domain |
| Pfam | PF02373 | 3.9E-36 | 222 | 341 | IPR003347 | JmjC domain |
| SMART | SM00355 | 9.7 | 1295 | 1317 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.887 | 1318 | 1347 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 15 | 1318 | 1342 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1320 | 1342 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 2.65E-9 | 1334 | 1376 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.2E-9 | 1345 | 1372 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.344 | 1348 | 1377 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.002 | 1348 | 1372 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1350 | 1372 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 4.42E-8 | 1366 | 1400 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.0E-9 | 1373 | 1401 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.284 | 1378 | 1409 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 1.1 | 1378 | 1404 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1380 | 1404 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1415 aa Download sequence Send to blast |
MSPPAVETPE WLRNLPVAPE YHPTAAEFVD PIAYILKIEA EASRYGICKI VPPLAPPPRE 60 ATVERLKASF AANAAAAAGV DGAAPAPTFP TRLQQVGFST KNRRPASRRV WESGERYTLE 120 AFRAKARDIE LPRHAVPPKH ATQLQLEALF WGACAARPFN VEYGNDMPGS GFAAPKEGGG 180 GNAALAARDV GETEWNMRLA PRARGSLLRA MGRDVAGVTT PMLYVAMLYS WFAWHVEDHE 240 LHSLNYLHFG KPKTWYGVPR DAMLAFEDAV RVHGYADDLN AIMAFQTLNE KTTVLSPEVL 300 LSAGVPCCRL VQNPGEFIIT FPGAYHSGFS HGFNCGEATN IATPRWLQVA KEAAIRRAST 360 NCGPLVSHYQ LLYELALSLR PRELKDSHGV PRSSRLRDKK KNEGEIMIKE TFVGSVIENN 420 NFLSILLDKS SCVIIPEIKF PLPSFPTSMA PEVTVKQGLI AGTCSISQKK AEDMLAYGNA 480 IDEIKGFEKM SESQSSSATT SSACNGRKLY ETKFGMVNSS ALLLNSEIQS GVIEKGGSHQ 540 GCGLLDQGRL PCVQCGILSY ACVAIIQPKE AAVQYVISQE CMSSSAKHGE IMKSNDTSNW 600 ITTVPPQGHS SETDDNRIHN MSSARVSDRC RQLYTSSTHG CASALGLLAS AYESSDSDEE 660 AEAPDNISNN SANNDAVNGI TNIQSSGTSV QYQNTNLHLY EEGCDSRATV SPMKPVENMS 720 IAMTQASIET DMTHLADLGE SLTAYDQWST YLDLDDDPTA PGVKASLSTS FSRAKGAMEP 780 DALTLLKYSK DSCRMHVFCL EHALETWTRL QQFGGANIVL LCHPEYPKAE SAAKVIAEEL 840 GMKHAWKDIT FKNATDEDIG RIRLALQDED AEPTSSDWAV KMGINIYYSA KQSKSPLYSK 900 QVPYNSIIYK AFAQENPDSL TDERQRSGTT KKKVAGSWCG KIWMSNQVHP LLDREREEQD 960 HDMVYSKAMV SVNSYDRIQE PSARSTSLIN RSLSKRVSRR KEGDSVERAK KKRYTASDVA 1020 TFDQPRNFDD HGKHESDGDE NENEDAQKTL QHQHESEKTN KNSSSKRRKD DKRNNNFYER 1080 HNDSDDHDYR LGIDSTENTT VGDCDNSPPQ GLDVVKVKSD TQLQGSKKKS SKCKASDDLS 1140 NVEKRLQKMV KKVSTKKHKN DKTNRQFQEK HSKDNNVDLL HEDNGDEATQ ENWDGVQQKT 1200 NDVKVKSRGK MHSGKKKASK CQTSDGLHNG DNEAKFSCDT DVCHRDKATI DKWEEIPKEK 1260 ADDVKVKSKM QSGKKKASKH PASDGLRNGD KGAKFSCDIE GCDMSFSTQQ DLALHKRDIC 1320 PVKGCKKKFF CHKYLLQHRK VHLDERPLMC SFTGCKKTFK WPWARTEHMR VHTGVRPYAC 1380 TEPGCTQTFR FVSDFSRHKR KTGHSCDKKK KNST* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 3e-68 | 10 | 379 | 7 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 3e-68 | 10 | 379 | 7 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 998 | 1012 | RRKEGDSVERAKKKR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
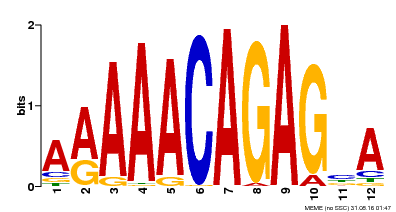
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sevir.3G311800.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025807104.1 | 0.0 | lysine-specific demethylase JMJ705-like | ||||
| TrEMBL | A0A2T7EFS1 | 0.0 | A0A2T7EFS1_9POAL; Uncharacterized protein | ||||
| STRING | BRADI4G39130.1 | 0.0 | (Brachypodium distachyon) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6718 | 30 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 1e-159 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sevir.3G311800.1.p |



