 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sevir.1G186800.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | M-type_MADS | ||||||||
| Protein Properties | Length: 255aa MW: 27143.2 Da PI: 8.0689 | ||||||||
| Description | M-type_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 55 | 1e-17 | 11 | 50 | 3 | 42 |
--SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-T CS
SRF-TF 3 ienksnrqvtfskRrngilKKAeELSvLCdaevaviifss 42
i n+ r tf kRr+g++KKA+EL +LC+++++v++++
Sevir.1G186800.1.p 11 IANDATRRATFKKRRKGLMKKASELATLCGVRACVVVYGA 50
8899**********************************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00432 | 7.7E-22 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 3.92E-23 | 1 | 86 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 17.577 | 1 | 49 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00266 | 2.17E-33 | 2 | 86 | No hit | No description |
| PRINTS | PR00404 | 1.1E-8 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 5.5E-18 | 11 | 49 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.1E-8 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0043078 | Cellular Component | polar nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 255 aa Download sequence Send to blast |
MARKKVALQW IANDATRRAT FKKRRKGLMK KASELATLCG VRACVVVYGA GESQPEVWPE 60 APGAAEDVVA RFRAVPELDQ CKKMLDMEGY LKQQVDKLRE QLHKAQRENR ERETALLLHD 120 AIAGRLPGLA GLSVEEVGSL GWMVENRIQA VRAAIAQLQG EGQDLPAAAL QLPQPSLPVA 180 PYYGIGAGAG HGEMMMQAPH PQGWMMAGGE IGALAYGGFV GTSTSAGADM PPQFGSMGAG 240 FAWPDTAGQS FPSM* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 2 | 24 | RKKVALQWIANDATRRATFKKRR |
| 2 | 21 | 25 | KKRRK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
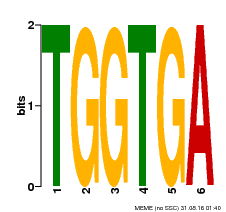
| Motif ID | Method | Source | Motif file |
| MP00553 | DAP | Transfer from AT5G48670 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sevir.1G186800.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KJ727526 | 1e-122 | KJ727526.1 Zea mays clone pUT5386 MADS transcription factor (MADS66) gene, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004954970.1 | 0.0 | agamous-like MADS-box protein AGL80 | ||||
| TrEMBL | A0A368PNV8 | 0.0 | A0A368PNV8_SETIT; Uncharacterized protein | ||||
| STRING | Si019470m | 1e-176 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP363 | 36 | 202 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G48670.1 | 1e-35 | AGAMOUS-like 80 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sevir.1G186800.1.p |



