 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Sevir.1G116900.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 424aa MW: 48375.1 Da PI: 8.2424 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.7 | 1e-05 | 115 | 137 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++Cp Cg +F + +Lk+H+++H
Sevir.1G116900.1.p 115 HRCPECGACFQKPAHLKQHMQSH 137
79*******************99 PP
| |||||||
| 2 | zf-C2H2 | 23.6 | 1.3e-07 | 143 | 167 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ Cp dC+ s++rk++L+rH+ +H
Sevir.1G116900.1.p 143 FICPleDCPFSYKRKDHLNRHMLKH 167
78*********************99 PP
| |||||||
| 3 | zf-C2H2 | 12.4 | 0.00047 | 174 | 197 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirt.H 23
C+ C+++Fs k n++rH + H
Sevir.1G116900.1.p 174 CTvdGCDRRFSIKANMQRHVKEfH 197
88889***************9988 PP
| |||||||
| 4 | zf-C2H2 | 16 | 3.4e-05 | 209 | 233 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
+ C+ C+k F+ s L++H +H
Sevir.1G116900.1.p 209 FICKeeGCNKAFKYLSKLRKHEESH 233
789999***************9988 PP
| |||||||
| 5 | zf-C2H2 | 20.6 | 1.2e-06 | 300 | 324 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirt.H 23
kC+ C +sFs+ksnL++H++ H
Sevir.1G116900.1.p 300 KCTfeGCERSFSNKSNLTKHMKAcH 324
79999*****************955 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00355 | 22 | 69 | 91 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 5.2E-5 | 112 | 134 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.002 | 115 | 137 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.006 | 115 | 142 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 117 | 137 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 5.91E-11 | 127 | 169 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 7.6E-12 | 135 | 169 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 1.9E-4 | 143 | 167 | IPR015880 | Zinc finger, C2H2-like |
| Pfam | PF00096 | 2.5E-5 | 143 | 167 | IPR007087 | Zinc finger, C2H2 |
| PROSITE profile | PS50157 | 9.037 | 143 | 172 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 145 | 167 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.1E-7 | 153 | 202 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.3E-8 | 170 | 199 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.512 | 172 | 202 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.027 | 172 | 197 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 174 | 197 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 4.3E-6 | 207 | 233 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0078 | 209 | 233 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.221 | 209 | 233 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 211 | 233 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 8.7 | 241 | 266 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 243 | 266 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 18 | 269 | 290 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 1.14E-8 | 284 | 341 | No hit | No description |
| PROSITE profile | PS50157 | 10.97 | 299 | 329 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.002 | 299 | 324 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.5E-8 | 300 | 324 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 301 | 324 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.0E-4 | 325 | 352 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 8.995 | 330 | 356 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 3.8 | 330 | 356 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 332 | 356 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 424 aa Download sequence Send to blast |
MKESLPPLSH SPPPPPGSGR SPLHSHSSCK LQARRTPVCS DSKSEMCSGD EIDGDASVGQ 60 KGFKDIRRYK CEFCTVVRSK KCLIQAHMVA HHKDELDKSE TYNSNGEKII CEEEHRCPEC 120 GACFQKPAHL KQHMQSHSNE RLFICPLEDC PFSYKRKDHL NRHMLKHQGK LFGCTVDGCD 180 RRFSIKANMQ RHVKEFHEDE NVTKSSQEFI CKEEGCNKAF KYLSKLRKHE ESHVKLDYVE 240 VVCCEPGCMK MFTNVECLKA HNQSCHQHVQ CEICGEKHLK KNIKRHLQAH DEVPTGARMK 300 CTFEGCERSF SNKSNLTKHM KACHDQLKLF ICRVAGCGKA FTYKHVRDNH EKSSAHVYVE 360 GDFEEMDEQL RSRPRGGRKR KALTVETLTR KRVTIPGEAS SLDDGEEYLR WLLSGGDDLG 420 QAQ* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 2e-18 | 119 | 324 | 19 | 188 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 2e-18 | 119 | 324 | 19 | 188 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 372 | 380 | RPRGGRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
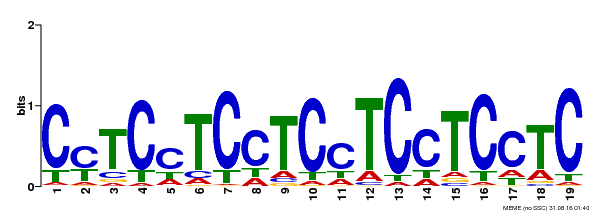
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Sevir.1G116900.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004952159.1 | 0.0 | transcription factor IIIA isoform X2 | ||||
| Swissprot | Q84MZ4 | 1e-117 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | A0A368PJD5 | 0.0 | A0A368PJD5_SETIT; Uncharacterized protein | ||||
| STRING | Si020193m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2382 | 38 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72050.2 | 1e-120 | transcription factor IIIA | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Sevir.1G116900.1.p |



