 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Seita.9G510100.2.p | ||||||||
| Common Name | SETIT_034076mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1024aa MW: 113950 Da PI: 5.2183 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 178.1 | 1.1e-55 | 9 | 125 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenp 92
ke ++rwl++ ei++iL+n++++++t e+++rp+sgsl+L++rk++ryfrkDG++w+kk+dgktv+E+he+LK g+++vl+cyYah+een+
Seita.9G510100.2.p 9 KEaQNRWLRPAEICEILKNYRNFRITPEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHERLKSGSIDVLHCYYAHGEENE 99
5559*************************************************************************************** PP
CG-1 93 tfqrrcywlLeeelekivlvhylevk 118
+fqrr+yw+Lee++ +ivlvhylevk
Seita.9G510100.2.p 100 NFQRRSYWMLEEDFMHIVLVHYLEVK 125
***********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 83.172 | 4 | 130 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.1E-77 | 7 | 125 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.3E-49 | 10 | 124 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 4.8E-8 | 434 | 519 | IPR013783 | Immunoglobulin-like fold |
| CDD | cd00102 | 2.57E-5 | 434 | 521 | No hit | No description |
| SuperFamily | SSF81296 | 1.03E-18 | 435 | 520 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 5.5E-9 | 437 | 514 | IPR002909 | IPT domain |
| CDD | cd00204 | 1.61E-12 | 615 | 727 | No hit | No description |
| SuperFamily | SSF48403 | 4.82E-16 | 626 | 730 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.2E-16 | 629 | 733 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 17.131 | 635 | 739 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.576 | 635 | 667 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.736 | 668 | 700 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.013 | 668 | 697 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 3500 | 707 | 736 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.89E-8 | 840 | 892 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.07 | 841 | 863 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.114 | 842 | 871 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0029 | 843 | 862 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0019 | 864 | 886 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.542 | 865 | 889 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.5E-4 | 867 | 886 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1024 aa Download sequence Send to blast |
MEDIEQILKE AQNRWLRPAE ICEILKNYRN FRITPEPPNR PPSGSLFLFD RKVLRYFRKD 60 GHNWRKKRDG KTVKEAHERL KSGSIDVLHC YYAHGEENEN FQRRSYWMLE EDFMHIVLVH 120 YLEVKGGKSS SRIRGHDDML QAARTDSPLS QLPSQTTEGE SSLSGQASEY EETESDIYSG 180 GAGYHPFSWM QQHENGGGPV MGASILSSYI PAPSVGNHQG FLATTTTTDF YSHGQDALPV 240 ALNEPGFGIA FDEADNQLDP SSLNGLVKPD QGVHRMAPPQ IADPSKQFPF TEGSGIESFT 300 FDEVYSNGLS IKDADTVGTD EESLWQLPGA ISSFPTEDSS QQNGRSLEEA INHPLLKTQS 360 SSLSDILKDS FKKNDSFTRW MSKELGEVDD SPIKSSSGVY WNSEDTDNII EASSRDQLDQ 420 FTVDPVVAQD QLFSIFDFSP SWAYAGSKTR VLITGRFLNS DEVQRCKWSC MFGEVEVPAE 480 ISADGTLRCY SPSHKPGRVP FYVTCSNRLA CSEIREFEFR PSNSQHIDGP TPHDIANKTY 540 LQMRLDDLLS LGQDEYQATV SNPTKDMIDL SKKISSLMTD NDSWSELLKL AGDNELATDD 600 KQDQFFENRV KEKLHIWLVH KAGDGGKGPS VLDEEGQGVL HLAAALGYDW AIRPTISAGV 660 SINFRDAHGW TALHWAAFCG RERTVVALIA LGAAPGALTD PTPDFPTGST PADLASANGY 720 KGISGFLAES SLTSHLQTLN LKEAMWSNAP EISGLPGIGD VTERKLSPLA GEGLLAGSMG 780 DSLGAVRNAT QAAARIYQVF RMQSFQRKQV VQYEDDNGAI SDDCALSLLS VKPSKPGQLD 840 PLHAAATRIQ NKYRGWKGRK EFLLIRQRII KIQAHVRGHQ VRKHYRKIIW SVGIVEKVIL 900 RWRRRGAGLR GFRSTEGAAE GTSSSGSDLI QNKPAEDDYD FLQQGRKQTE ERLQKALARV 960 KSMVQYPDAR DQYQRILNVV TKMQESQAMQ EKMLESSTDM DEGLAMSDFE ELWGDDMPMP 1020 GYI* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
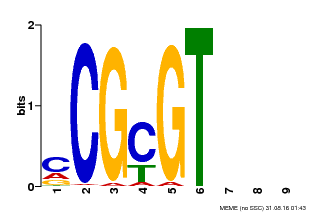
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Seita.9G510100.2.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004985393.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X2 | ||||
| TrEMBL | A0A368SUS5 | 0.0 | A0A368SUS5_SETIT; Uncharacterized protein | ||||
| STRING | Si034076m | 0.0 | (Setaria italica) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Seita.9G510100.2.p |



