 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Seita.6G247000.1.p | ||||||||
| Common Name | LOC101765464, SETIT_013546mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Cenchrinae; Setaria
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 530aa MW: 54704.1 Da PI: 9.7927 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 17.4 | 1.3e-05 | 66 | 88 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
++C+ C+k F r nL+ H r H
Seita.6G247000.1.p 66 FVCEVCNKGFQREQNLQLHRRGH 88
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 13.2 | 0.00026 | 142 | 164 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC +C+k++ +s+ k H +t+
Seita.6G247000.1.p 142 WKCDKCNKRYAVQSDWKAHSKTC 164
58*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 1.5E-7 | 65 | 88 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 7.2E-6 | 65 | 88 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0041 | 66 | 88 | IPR015880 | Zinc finger, C2H2-like |
| Pfam | PF12171 | 3.4E-5 | 66 | 88 | IPR022755 | Zinc finger, double-stranded RNA binding |
| PROSITE profile | PS50157 | 11.094 | 66 | 88 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 68 | 88 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 300 | 107 | 137 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 3.9E-5 | 130 | 163 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 1.5E-7 | 137 | 162 | No hit | No description |
| SMART | SM00355 | 110 | 142 | 162 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 530 aa Download sequence Send to blast |
MAASAHLFGL GDAQMQPRPP QQQAAPPPPA APPAPKKKRN QPGNPKDPDA EVIALSPKTL 60 LATNRFVCEV CNKGFQREQN LQLHRRGHNL PWKLKQKNPK ETRRRVYLCP EPTCVHHDPS 120 RALGDLTGIK KHYCRKHGEK KWKCDKCNKR YAVQSDWKAH SKTCGTREYR CDCGTLFSRR 180 DSFITHRAFC DALAQENARV PPIGAGMYGA GGMAFGLSGM AASQLQSFQD QAHSSATTAI 240 SGNPAAQFEH LMPTSTSFRG AQPASSSSSP FYLGGAEDGN QSQPGHTSLL HGKPAFHGLM 300 QLPEQHGQPG SNGLLNLGFF SGASSGGQDA RLVFPGQFNG AAGGNGRGDG GEHGNSSANT 360 ESAAIFSGNL MGNQMTGGGG GGFSSSLYNS AGTVAPPQMS ATALLQKAAQ MGATSSGGGG 420 GSANSLLKGL GSGGALNGRA AGAAGFMAGE SSSRSASQAE NESQFRDLMN SLAASGSSGF 480 PGLDDGKPST RDFLGVGGGV VRSMGGAAGL PLRHGAAGIG MGSLDPEMK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 6e-34 | 138 | 200 | 3 | 65 | Zinc finger protein JACKDAW |
| 5b3h_F | 6e-34 | 138 | 200 | 3 | 65 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor acting as a positive regulator of the starch synthase SS4. Controls chloroplast development and starch granule formation (PubMed:22898356). Binds DNA via its zinc fingers (PubMed:24821766). Recognizes and binds to SCL3 promoter sequence 5'-AGACAA-3' to promotes its expression when in complex with RGA (PubMed:24821766). {ECO:0000269|PubMed:22898356, ECO:0000269|PubMed:24821766}. | |||||
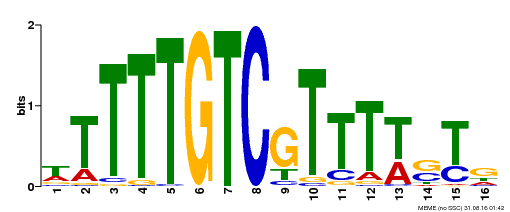
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00255 | DAP | Transfer from AT2G02070 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Seita.6G247000.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004974170.1 | 0.0 | protein indeterminate-domain 5, chloroplastic isoform X1 | ||||
| Swissprot | Q9ZUL3 | 1e-120 | IDD5_ARATH; Protein indeterminate-domain 5, chloroplastic | ||||
| TrEMBL | K3YH28 | 0.0 | K3YH28_SETIT; Uncharacterized protein | ||||
| STRING | Si013546m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP202 | 38 | 320 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G02080.1 | 1e-111 | indeterminate(ID)-domain 4 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Seita.6G247000.1.p |
| Entrez Gene | 101765464 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



